 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | CCG020422.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Malpighiales; Salicaceae; Saliceae; Populus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 362aa MW: 40653.4 Da PI: 9.2753 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.9 | 4e-32 | 70 | 123 | 2 | 55 |
G2-like 2 prlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
prlrWtp+LH +Fv+ave+LGG+e+AtPk +l++m++kgL+++hvkSHLQ+YR+
CCG020422.1 70 PRLRWTPDLHLCFVHAVERLGGQERATPKLVLQMMNIKGLSIAHVKSHLQMYRS 123
8****************************************************7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.6E-30 | 66 | 125 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.096 | 66 | 126 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.06E-14 | 68 | 125 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.5E-22 | 70 | 124 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.6E-9 | 71 | 122 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 362 aa Download sequence Send to blast |
MEGSDSFGGS KSSYSNKEVE QDEGDEEEVD KISFKSRNEG VSTSNSSVEE NEKKAASTGS 60 VRQYIRSKMP RLRWTPDLHL CFVHAVERLG GQERATPKLV LQMMNIKGLS IAHVKSHLQM 120 YRSKRSDEPN QGQGLFLEGG DRLIYNHSQL PVLQNFNQRS SCNLRYGNAS WRGHDHQMYS 180 PYKEGTALNR FKHGLYGSVS ERLVIGRNNH NSLNYDSSIN IPSLNVQATS RTHQFLEGVK 240 LFQVSRREES RPGSMESNFI AKLQERSGID QKECLNTTSS ADKNWRTIQE MQKGSKRKTS 300 DSDCNLELNL ALKLSTKDED GLQKCVADGS LSLSLSSKLG RSMEGDGSRK HARMASTLHL 360 TL |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 9e-18 | 71 | 125 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 9e-18 | 71 | 125 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 9e-18 | 71 | 125 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 9e-18 | 71 | 125 | 3 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
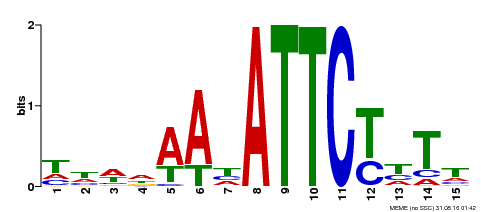
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00301 | DAP | Transfer from AT2G40260 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011035900.1 | 0.0 | PREDICTED: putative two-component response regulator ARR21 | ||||
| TrEMBL | B9HHT8 | 0.0 | B9HHT8_POPTR; Uncharacterized protein | ||||
| STRING | POPTR_0010s19310.1 | 0.0 | (Populus trichocarpa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF2378 | 34 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G40260.1 | 6e-53 | G2-like family protein | ||||



