 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_27747 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | RAV | ||||||||
| Protein Properties | Length: 322aa MW: 35709.6 Da PI: 9.9231 | ||||||||
| Description | RAV family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | AP2 | 39.6 | 1.3e-12 | 55 | 103 | 1 | 55 |
AP2 1 sgykGVrwdkkrgrWvAeIrdpsengkrkrfslgkfgtaeeAakaaiaarkkleg 55
s+ykGV + +grW A+I++ ++r++lg+f + +Aa+a++ a+++++g
PEQU_27747 55 SQYKGVVPQP-NGRWGAQIYEK-----HRRVWLGTFNLEHDAARAYDLAAQRFRG 103
78****9888.8*********3.....5**********99*************98 PP
| |||||||
| 2 | B3 | 90.3 | 1.5e-28 | 171 | 271 | 1 | 97 |
EEEE-..-HHHHTT-EE--HHH.HTT....---..--SEEEEEE..TTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE..E CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh....ggkkeesktltled..esgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsefelv 93
f+k+ tpsdv+k++rlv+pk++ae+h +g +++ l +e g++W+++++y+++s++yvltkGW++Fvk++gL +gD v F++++ +e++++
PEQU_27747 171 FDKAVTPSDVGKLNRLVIPKQHAEKHfplaDGA--KGVLLNFEEgcAGGKVWRFRYSYWNSSQSYVLTKGWSRFVKEKGLVAGDLVGFYRSKGEERQFY 267
89************************9975333..66677777766666****************************************8877777677 PP
EEEE CS
B3 94 vkvf 97
+ +
PEQU_27747 268 IDCR 271
7665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF54171 | 5.69E-17 | 55 | 111 | IPR016177 | DNA-binding domain |
| CDD | cd00018 | 7.90E-26 | 55 | 111 | No hit | No description |
| Gene3D | G3DSA:3.30.730.10 | 1.7E-20 | 56 | 111 | IPR001471 | AP2/ERF domain |
| Pfam | PF00847 | 1.4E-8 | 56 | 103 | IPR001471 | AP2/ERF domain |
| SMART | SM00380 | 2.6E-28 | 56 | 117 | IPR001471 | AP2/ERF domain |
| PROSITE profile | PS51032 | 20.35 | 56 | 111 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.1E-7 | 57 | 68 | IPR001471 | AP2/ERF domain |
| PRINTS | PR00367 | 9.1E-7 | 93 | 113 | IPR001471 | AP2/ERF domain |
| Gene3D | G3DSA:2.40.330.10 | 2.3E-38 | 162 | 276 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 1.7E-28 | 168 | 273 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.68E-27 | 169 | 264 | No hit | No description |
| SMART | SM01019 | 9.1E-20 | 171 | 273 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 13.719 | 171 | 272 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.1E-25 | 171 | 271 | IPR003340 | B3 DNA binding domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009910 | Biological Process | negative regulation of flower development | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048366 | Biological Process | leaf development | ||||
| GO:0048527 | Biological Process | lateral root development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 322 aa Download sequence Send to blast |
MQSNSCNSSS SSTLDLPSPT LHPIHSNHSA ITSPAPTTPS SEPSTEAESK KLPSSQYKGV 60 VPQPNGRWGA QIYEKHRRVW LGTFNLEHDA ARAYDLAAQR FRGRDAITNF PPLSLSADPH 120 ELSFLLSRSK AEIVDMLRKH TYLDELRLCP PLFSPKNASV VSSKPAREHL FDKAVTPSDV 180 GKLNRLVIPK QHAEKHFPLA DGAKGVLLNF EEGCAGGKVW RFRYSYWNSS QSYVLTKGWS 240 RFVKEKGLVA GDLVGFYRSK GEERQFYIDC RPRGEEGGGV QTFRLFGVNI GILGSLNMNK 300 KRSFEGMEVG LLMKKQCVGG LG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1wid_A | 5e-48 | 168 | 273 | 11 | 117 | DNA-binding protein RAV1 |
| Search in ModeBase | ||||||
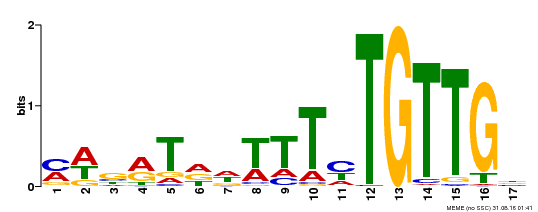
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00139 | DAP | Transfer from AT1G13260 | Download |
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020579562.1 | 0.0 | AP2/ERF and B3 domain-containing protein Os05g0549800-like | ||||
| Swissprot | Q6L4H4 | 1e-109 | Y5498_ORYSJ; AP2/ERF and B3 domain-containing protein Os05g0549800 | ||||
| TrEMBL | A0A2I0W076 | 0.0 | A0A2I0W076_9ASPA; AP2/ERF and B3 domain-containing protein | ||||
| STRING | XP_008799316.1 | 1e-116 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP456 | 38 | 203 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G13260.1 | 1e-104 | related to ABI3/VP1 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



