| Signature Domain? help Back to Top |
 |
| No. |
Domain |
Score |
E-value |
Start |
End |
HMM Start |
HMM End |
| 1 | SRF-TF | 87.8 | 5.9e-28 | 9 | 59 | 1 | 51 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE- CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeyss 51
k+ien +nrqvt+skRr+gi+KKA E+ vLCdaev++i+fsstgk+ ey+s
PEQU_18912 9 KKIENPTNRQVTYSKRRAGIMKKAREITVLCDAEVSLIMFSSTGKFSEYCS 59
68***********************************************96 PP
|
| 2 | K-box | 82.6 | 9.1e-28 | 71 | 169 | 1 | 99 |
K-box 1 yqkssgksleeakaeslqqelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkkle 99
yq+ sg +l+++++e++ + l+ k+ ++nL+re R+++GedLe L++keL+ Leq+++++lk +R++K++++ +q+++ +kk k+ qe++++L ++le
PEQU_18912 71 YQQVSGINLWSSQYEKMLNTLNHSKEINRNLRREVRQRMGEDLEGLDIKELRGLEQNIDEALKLVRNRKYHVISTQTDTYKKKLKNSQETHRNLMHELE 169
7899999***************************************************************************************99986 PP
|
| Protein Features
? help Back to Top |
 |
| Database |
Entry ID |
E-value |
Start |
End |
InterPro ID |
Description |
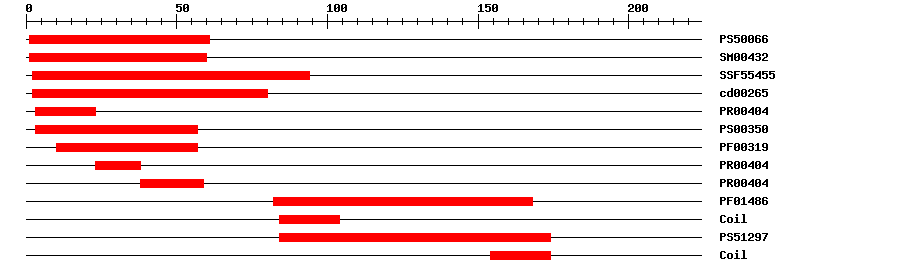
| PROSITE profile | PS50066 | 31.511 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 2.0E-40 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| SuperFamily | SSF55455 | 6.02E-37 | 2 | 94 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 3.08E-41 | 2 | 80 | No hit | No description |
| PRINTS | PR00404 | 2.8E-27 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 1.4E-22 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.8E-27 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 2.8E-27 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF01486 | 1.2E-17 | 82 | 168 | IPR002487 | Transcription factor, K-box |
| PROSITE profile | PS51297 | 15.027 | 84 | 174 | IPR002487 | Transcription factor, K-box |
| Publications
? help Back to Top |
- Kikuchi S, et al.
Collection, mapping, and annotation of over 28,000 cDNA clones from japonica rice.
Science, 2003. 301(5631): p. 376-9
[PMID:12869764] - Tsai WC,Kuoh CS,Chuang MH,Chen WH,Chen HH
Four DEF-like MADS box genes displayed distinct floral morphogenetic roles in Phalaenopsis orchid.
Plant Cell Physiol., 2004. 45(7): p. 831-44
[PMID:15295066] - Chen ZX, et al.
Morphogenesis and molecular basis on naked seed rice, a novel homeotic mutation of OsMADS1 regulating transcript level of AP3 homologue in rice.
Planta, 2006. 223(5): p. 882-90
[PMID:16254725] - Zhang Q, et al.
Morphological, anatomical and genetic analysis for a rice mutant with abnormal hull.
J Genet Genomics, 2007. 34(6): p. 519-26
[PMID:17601611] - Yoshida H, et al.
superwoman1-cleistogamy, a hopeful allele for gene containment in GM rice.
Plant Biotechnol. J., 2007. 5(6): p. 835-46
[PMID:17764519] - Yao SG,Ohmori S,Kimizu M,Yoshida H
Unequal genetic redundancy of rice PISTILLATA orthologs, OsMADS2 and OsMADS4, in lodicule and stamen development.
Plant Cell Physiol., 2008. 49(5): p. 853-7
[PMID:18378529] - Xiao H, et al.
STAMENLESS 1, encoding a single C2H2 zinc finger protein, regulates floral organ identity in rice.
Plant J., 2009. 59(5): p. 789-801
[PMID:19453444] - Seok HY, et al.
Rice ternary MADS protein complexes containing class B MADS heterodimer.
Biochem. Biophys. Res. Commun., 2010. 401(4): p. 598-604
[PMID:20888318] - Li H, et al.
Rice MADS6 interacts with the floral homeotic genes SUPERWOMAN1, MADS3, MADS58, MADS13, and DROOPING LEAF in specifying floral organ identities and meristem fate.
Plant Cell, 2011. 23(7): p. 2536-52
[PMID:21784949] - Sato H,Yoshida K,Mitsuda N,Ohme-Takagi M,Takamizo T
Male-sterile and cleistogamous phenotypes in tall fescue induced by chimeric repressors of SUPERWOMAN1 and OsMADS58.
Plant Sci., 2012. 183: p. 183-9
[PMID:22195592] - Ohmori S,Tabuchi H,Yatou O,Yoshida H
Agronomic traits and gene containment capability of cleistogamous rice lines with the superwoman1-cleistogamy mutation.
Breed. Sci., 2012. 62(2): p. 124-32
[PMID:23136523] - Yun D, et al.
OsMADS16 genetically interacts with OsMADS3 and OsMADS58 in specifying floral patterning in rice.
Mol Plant, 2013. 6(3): p. 743-56
[PMID:23300256] - Hsu CC, et al.
Histone acetylation accompanied with promoter sequences displaying differential expression profiles of B-class MADS-box genes for phalaenopsis floral morphogenesis.
PLoS ONE, 2014. 9(12): p. e106033
[PMID:25501842] - Lombardo F, et al.
The superwoman1-cleistogamy2 mutant is a novel resource for gene containment in rice.
Plant Biotechnol. J., 2017. 15(1): p. 97-106
[PMID:27336225]
|





