 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_16477 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 962aa MW: 107112 Da PI: 6.9922 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 119.1 | 2.2e-37 | 145 | 219 | 2 | 76 |
-SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S- CS
SBP 2 CqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkk 76
Cqv++C a+ls+ k+yhrrhkvCe+h+ka++++v ++ qrfCqqCsrfh l+efDe+krsCrrrLa+hn+rrrk+
PEQU_16477 145 CQVQSCCANLSQSKDYHRRHKVCETHAKASSAVVGNVIQRFCQQCSRFHLLQEFDEGKRSCRRRLAGHNRRRRKT 219
*************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 1.7E-30 | 138 | 206 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.428 | 142 | 219 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 7.98E-35 | 143 | 220 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 6.5E-28 | 145 | 218 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 6.3E-9 | 709 | 844 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 2.92E-12 | 711 | 841 | No hit | No description |
| SuperFamily | SSF48403 | 1.55E-9 | 737 | 843 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 962 aa Download sequence Send to blast |
METNFGGDNR QFFAAGSSKR FGKEAQEWDL NDWKWDGDLF MATPVNPGPL NGLNKNLFPD 60 IVLSNSSSSC SDETEFGLGN VLGEADKRRR IVVAEDDEQC DDTGSLTLKL GAHAYPVMEE 120 DFSLPGMKNG KRVMMQASNS NHPKCQVQSC CANLSQSKDY HRRHKVCETH AKASSAVVGN 180 VIQRFCQQCS RFHLLQEFDE GKRSCRRRLA GHNRRRRKTL SGAATPSESS AAGDQSAGYL 240 LISLLRILAN LNSDRSEQSN DQELLSNFLR NIVSLAGSSD GKNSSGLLQA SSDLQKIGTS 300 AGTSGEAANA LISACFDETR QATCSKRLVP EKNCSPDKSP TVDSLQHMLP TACTVASGRE 360 KAFDLNNVYY EEEDGALGCG QSAKQATLDS GYANCPWMLQ DSHQSSPPQA SGNTDSTSNR 420 SPSSSNGEAQ SRTDRIVFKL FGKEPNDFPL VLRAQIFDWL SNSPTDMESY IRPGCIVLTV 480 YLCLEVSMWD ELCHDLGSSL KRLLFLSNDD FWRIGWIYAR VQHHIVFIYN GQVVLNKSLI 540 MERSDCNRIV AVTPIAVPPS SKVNFKVKGY NLVRSTTRLL CSFEGKYLIK ETDEPTIQYS 600 DYSSSGNRNE QLQSLSFSCH IPDTNGRGFI EIMQLQRLNP CFFPFIVAEE DICAEIQVLE 660 ISEELSEKNL NFLHEIGWLL RRSQLRSVQK LSLSRFGPIL RFTIDHDWCA IVKKLLDILF 720 EGNIETYGRS PIDLLMSEYL LHYAVQRNSN LTVKLLLGYK LDKISDGGLN NLFRPDVRGT 780 SGITPLHVAA TSSFAESMLN ILTDDPGQFG VKAWTNARDA TGFTPEDYAR ARGHESYILL 840 LQKKINKITE KCDVVVNMPC KLSLPSDAIY RQSEGLKPAK LHSLGIYNTK SKQSEPPFCK 900 LCEQHLACRS SVRGSLLYRP MLLSMVGIAA VCVCVGLLFK GPPEVMFVYP PFRWELLESG 960 YI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-29 | 145 | 218 | 11 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 86 | 90 | KRRRI |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
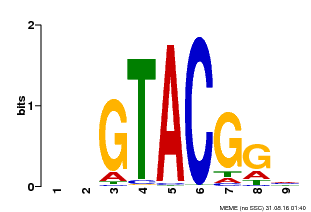
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020575747.1 | 0.0 | LOW QUALITY PROTEIN: squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A2I0V7E8 | 0.0 | A0A2I0V7E8_9ASPA; Squamosa promoter-binding-like protein 12 | ||||
| STRING | XP_008810336.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1998 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||



