 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_16180 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 678aa MW: 74221.6 Da PI: 6.3303 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 90.3 | 1.6e-28 | 219 | 272 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv+av+qL G++kA+Pk+ilelm+v+gLt+e+v+SHLQk+Rl
PEQU_16180 219 KPRVVWSVELHQQFVNAVNQL-GIDKAVPKRILELMNVPGLTRENVASHLQKFRL 272
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 78.6 | 2.2e-26 | 34 | 142 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTTSEEEEEESTTTHHHHHHHHHTTES CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepklpiivvtahgeeedalealkaGak 95
vl+vdD+ + +++l+ +l+k +y +v++++++ al ll+e++ +D+++ D+ mp+mdG++ll+ e++lp+i+++a g + + + +k Ga
PEQU_16180 34 VLVVDDDVTCLKILEHMLQKCRY-HVTTCSEALTALSLLRERKggFDVVISDVHMPDMDGFKLLELVGL-EMDLPVIMMSADGRTSAVMRGIKHGAC 128
89*********************.***************999999*******************87754.558************************ PP
EEEESS--HHHHHH CS
Response_reg 96 dflsKpfdpeelvk 109
d+l Kp+ +eel +
PEQU_16180 129 DYLIKPVRIEELKN 142
***********987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PIRSF | PIRSF036392 | 1.5E-167 | 11 | 636 | IPR017053 | Response regulator B-type, plant |
| SuperFamily | SSF52172 | 1.42E-36 | 31 | 161 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 9.2E-43 | 31 | 171 | No hit | No description |
| SMART | SM00448 | 5.5E-30 | 32 | 144 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 42.532 | 33 | 148 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 4.4E-23 | 34 | 142 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 4.18E-27 | 35 | 147 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.3E-30 | 216 | 277 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 11.002 | 216 | 275 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.34E-19 | 217 | 276 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.8E-25 | 219 | 272 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 6.4E-7 | 221 | 270 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 678 aa Download sequence Send to blast |
MQKLSAQSYS SCRADAGEST VLSSPELFPA GLRVLVVDDD VTCLKILEHM LQKCRYHVTT 60 CSEALTALSL LRERKGGFDV VISDVHMPDM DGFKLLELVG LEMDLPVIMM SADGRTSAVM 120 RGIKHGACDY LIKPVRIEEL KNIWQHVVRK KWNENRDVEH SGSFEDSDRQ RRLADEAEYD 180 SSINEGTDGS WKSQKKKRDS KDDEEDGELD NEDPSASKKP RVVWSVELHQ QFVNAVNQLG 240 IDKAVPKRIL ELMNVPGLTR ENVASHLQKF RLYLKRISGV TQHQSVLPNP FCAPVEPNTK 300 VGSVGRLDFH SLAATGQIPP QTLAVLHAEL LGRPSGNIVL PAIDSTVLLQ ASSQNSKFLP 360 SEKEIAFGQP LFKCQSGISK PCPPSGINTQ DTPSSFSSWP SSQVAITASN NELGGLNNAQ 420 NNHLLMQMLQ HQQQQQHHQE EPFNLVASES HNAINVQPSC LVVPSQLPSS LNGGNCSMVV 480 APQQSSNIKL GSSSVPTSNS TFNGTGITFD YTIHSSQSTN ATLGMGRTDS DFKNIGLSSS 540 FSPLASVSPS VSSSSVCTSV GGWRLQNSNM NFGFHKSLGQ VPNSRFLGPE VKSPAISVNY 600 ENVRNSGFVG KGTCLPSRFA VDDIESPTND WNNMKMSMFD DVDKVKQELN LDFLEGSKVE 660 KTMLPNFPSN DFMGVVSK |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 6e-23 | 215 | 278 | 1 | 64 | ARR10-B |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that binds specific DNA sequence. Functions as a response regulator involved in His-to-Asp phosphorelay signal transduction system. Phosphorylation of the Asp residue in the receiver domain activates the ability of the protein to promote the transcription of target genes. May directly activate some type-A response regulators in response to cytokinins. {ECO:0000250|UniProtKB:Q940D0}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
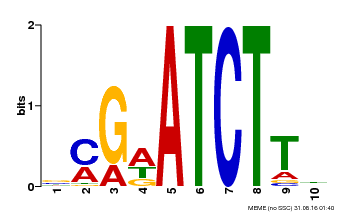
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | FN908141 | 2e-46 | FN908141.1 Populus x canadensis mRNA for putative B-type response regulator 16 (rr16 gene), cultivar Dorskamp. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020576239.1 | 0.0 | two-component response regulator ORR21-like isoform X2 | ||||
| Swissprot | Q8H7S7 | 1e-176 | ORR21_ORYSJ; Two-component response regulator ORR21 | ||||
| TrEMBL | A0A2I0VIL4 | 0.0 | A0A2I0VIL4_9ASPA; Two-component response regulator ARR2 | ||||
| STRING | XP_008812974.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5796 | 37 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 1e-119 | response regulator 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



