 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_16048 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1055aa MW: 120344 Da PI: 6.2574 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 179.9 | 3.2e-56 | 11 | 127 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvggvevlycyYahseenptfqrrcyw 100
+e ++rwl++ ei++iL n++k++++ e++++p+sgsl+L++rk++ryfrkDG++w+kkkdgktv+E+hekLK g+++vl+cyYah+e++++fqrr+yw
PEQU_16048 11 EEaQHRWLRPLEICEILRNYHKFHIAPEPPNKPPSGSLFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKAGSIDVLHCYYAHGEDDENFQRRSYW 109
4559*********************************************************************************************** PP
CG-1 101 lLeeelekivlvhylevk 118
+Leeel +ivlvhy evk
PEQU_16048 110 MLEEELMHIVLVHYHEVK 127
**************9985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 82.137 | 6 | 132 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 3.2E-78 | 9 | 127 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 4.8E-49 | 12 | 125 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 1.7E-5 | 492 | 579 | IPR013783 | Immunoglobulin-like fold |
| CDD | cd00102 | 0.00415 | 492 | 579 | No hit | No description |
| SuperFamily | SSF81296 | 4.2E-16 | 493 | 578 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 2.4E-7 | 493 | 578 | IPR002909 | IPT domain |
| SuperFamily | SSF48403 | 1.12E-18 | 675 | 788 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 9.6E-18 | 676 | 789 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 3.19E-15 | 688 | 786 | No hit | No description |
| PROSITE profile | PS50297 | 17.316 | 689 | 788 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 3600 | 694 | 723 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 8.603 | 694 | 726 | IPR002110 | Ankyrin repeat |
| Pfam | PF12796 | 5.8E-7 | 699 | 789 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 0.027 | 727 | 756 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 760 | 766 | 795 | IPR002110 | Ankyrin repeat |
| SMART | SM00015 | 0.043 | 900 | 922 | IPR000048 | IQ motif, EF-hand binding site |
| SuperFamily | SSF52540 | 2.04E-8 | 901 | 951 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| Pfam | PF00612 | 5.5E-4 | 902 | 921 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 8.736 | 902 | 930 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.011 | 923 | 945 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.871 | 924 | 948 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.5E-4 | 928 | 945 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1055 aa Download sequence Send to blast |
ISSIDLEQIL EEAQHRWLRP LEICEILRNY HKFHIAPEPP NKPPSGSLFL FDRKVLRYFR 60 KDGHNWRKKK DGKTVKEAHE KLKAGSIDVL HCYYAHGEDD ENFQRRSYWM LEEELMHIVL 120 VHYHEVKGNK TKFLQPREDG TISRITHVDS PARSNSSVNQ SQSPSLAANL ESPNITHLSE 180 YEDDSADIFQ SSSRYNSFLT MQQYGPRMNV KLANSCDIAP EKDHPCGYQR LQAEIPTFNS 240 FTVAQVGMHS LANESDGGLT FTGTKTMNDM ASWHEIFERC SMELESSNFD MIVASTCANT 300 MEEFPRQDSV ENILNSGLLD ANEDGAAVLS KPIWQVRHNF MHFMMKILLR WHTFSFSSQK 360 STHYVFIVPT KTRNYFLSLH DVNLQFYDDE VGSLIPSITE AENIQKSDTN HMNQLSNLSS 420 IDNQDLKKYD SFSRWASNEL GEVDDSLITL NSGLCWSTLE NDSVVAEPSV HNEEHFDTYL 480 MGPSISHDQL FSIIDFFPNW TYTELETKVL VKGKFLMNKE DVQKYSWSCM FGNIEVPAYI 540 LADGNLCCYT PSHKPGKIPF YVTRSNRFAC SEIREFEYKL NNMQKFENTD SDSSNTNDID 600 LLFRFEKLLV LENSDHTSPD SSHERPHSNC TICSMMMEVD VELLNLFKHA PDEDSPHIAA 660 ENWLLEEPLK EKLHVWLLHK VGEDVKGPNV LDNEGQGVLH LAAALGYDWA IKPIIAAGVG 720 INFRDVHGWT ALHWAAFFGR ERTVVTLIAL DASPGALTDP TPEFPSGRTP ADLASQNGYK 780 GIAGFLAESS LLKHLRELTL KDSKGCDLTE SSGLLDYHHS TDMGSSHNAF GGTQTSLKTS 840 LSAVRNATEA AARIYQVFRV QSFHRKKIMD YGDENCRISD EDAIPLISVK NNKSGQHGMP 900 VHAAAIKIQN KFRGWKGRKE FLTIRQQIVK IQAHVRGHQV RKHYRKILWS VGIVEKVILR 960 WRRKGSGLRG FRSDEPLEAR ISENQDMKED DYDFLQEGRK QTEARLEKAL ARVKSMVQYP 1020 EAIDQYRRLL NAVEELQESD VSSTFSLFTH IANNS |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
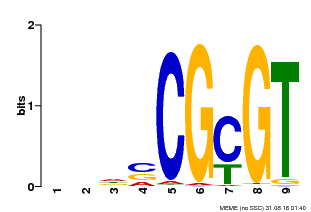
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020575860.1 | 0.0 | calmodulin-binding transcription activator 2 isoform X3 | ||||
| TrEMBL | A0A2I0WZX2 | 0.0 | A0A2I0WZX2_9ASPA; Calmodulin-binding transcription activator 3 | ||||
| STRING | XP_008795547.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP608 | 38 | 140 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||



