 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PEQU_08461 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; Asparagales; Orchidaceae; Epidendroideae; Vandeae; Aeridinae; Phalaenopsis
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 240aa MW: 28163.2 Da PI: 9.8348 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 109.1 | 3.3e-34 | 17 | 109 | 2 | 103 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellekik 98
F+ k+y++++d++++++isw +n+nsfvvld+ f +++Lp +Fkhsnf+SFvRQLn+YgF+kv+ ++ weF+h+sF +g+ +ll++i
PEQU_08461 17 FVAKIYQMVSDPANDSIISWGRNSNSFVVLDPFGFCQNLLPLHFKHSNFSSFVRQLNTYGFRKVNPDH---------WEFAHPSFLRGQSQLLSRIA 104
9*****************************************************************99.........******************** PP
XXXXX CS
HSF_DNA-bind 99 rkkse 103
rk++e
PEQU_08461 105 RKRKE 109
*9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00415 | 5.1E-51 | 13 | 106 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene3D | G3DSA:1.10.10.10 | 1.9E-36 | 14 | 106 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 1.56E-33 | 14 | 106 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| PRINTS | PR00056 | 2.1E-17 | 17 | 40 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 1.0E-29 | 17 | 106 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.1E-17 | 55 | 67 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 56 | 80 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.1E-17 | 68 | 80 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 240 aa Download sequence Send to blast |
MDIAGDCCGG EFAAPPFVAK IYQMVSDPAN DSIISWGRNS NSFVVLDPFG FCQNLLPLHF 60 KHSNFSSFVR QLNTYGFRKV NPDHWEFAHP SFLRGQSQLL SRIARKRKER DVEIDEDDRA 120 LVMELRRLKE EQRAIEDKLE RLSRRVMETE RRPRKMLQFL VRVFEDPQML GRLKGRAGIG 180 EGEGFGWEKK ERLRIDGEPP PPQPWERWFR VGFGGRWGRF WLGSVSYGFS LIFVFFFFSF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5d5u_B | 6e-26 | 1 | 106 | 12 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 6e-26 | 1 | 106 | 12 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 6e-26 | 1 | 106 | 12 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 104 | 128 | RKRKERDVEIDEDDRALVMELRRLK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00379 | DAP | Transfer from AT3G24520 | Download |
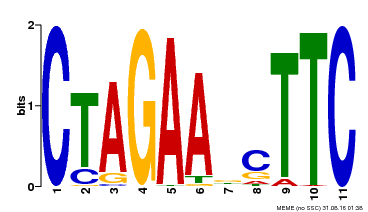
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_020587759.1 | 1e-175 | heat stress transcription factor C-2a-like | ||||
| Swissprot | Q6EUG4 | 2e-64 | HFC2A_ORYSJ; Heat stress transcription factor C-2a | ||||
| TrEMBL | A0A080YTS6 | 7e-69 | A0A080YTS6_WHEAT; Uncharacterized protein | ||||
| STRING | LPERR02G08320.1 | 3e-67 | (Leersia perrieri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4471 | 34 | 70 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24520.1 | 3e-60 | heat shock transcription factor C1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



