 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s987991g004 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 322aa MW: 36284.7 Da PI: 9.8755 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.9 | 2.2e-41 | 230 | 307 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+CqvegC++ l +ak+yh+rhkvCe+hskap+v+v g eqrfCqqCsrfh +sefD++krsCrrrLa+hnerrrk+++
PDK_30s987991g004 230 RCQVEGCNKALGDAKDYHKRHKVCEMHSKAPKVVVLGAEQRFCQQCSRFHVISEFDDAKRSCRRRLAGHNERRRKSTN 307
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 7.9E-35 | 225 | 292 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.265 | 228 | 305 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 5.76E-39 | 229 | 310 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 5.5E-32 | 231 | 304 | IPR004333 | Transcription factor, SBP-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010228 | Biological Process | vegetative to reproductive phase transition of meristem | ||||
| GO:0010229 | Biological Process | inflorescence development | ||||
| GO:0010321 | Biological Process | regulation of vegetative phase change | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 322 aa Download sequence Send to blast |
MASSNDSGSI ELPQQSRFQF ECDPLLGHKQ VANLEYLSTL GINQGGPALK EMDQELPAHV 60 IISSLHVKLD RHHAASIRGV VEILHELLSN EDVVHDASAT DECLFLWTIL GRKEKKAAYF 120 PTFHWGSLIN RVRRKEFGYC LSFKPARALR RGLRCLSRDP THLQPFLPPL LASRGSRLPV 180 CPLFRKESHH HHHLTCLKLG KRQYYGEEGV AGGGIKRERL SSNPAAGVPR CQVEGCNKAL 240 GDAKDYHKRH KVCEMHSKAP KVVVLGAEQR FCQQCSRFHV ISEFDDAKRS CRRRLAGHNE 300 RRRKSTNDSI KNSSLVQVLV LG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 1e-30 | 222 | 304 | 2 | 84 | squamosa promoter binding protein-like 4 |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00290 | DAP | Transfer from AT2G33810 | Download |
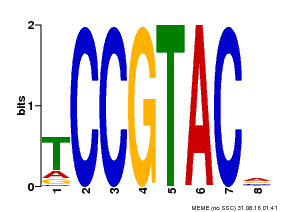
| |||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP10303 | 34 | 39 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G33810.1 | 4e-38 | squamosa promoter binding protein-like 3 | ||||



