 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s948451g002 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 332aa MW: 37130 Da PI: 6.6375 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 109.2 | 3e-34 | 19 | 111 | 2 | 103 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkel 93
F+ k+y++++d++++ +i+w + +nsf+vld+ +f++ +Lp+yFkhsnf+SFvRQLn+YgFkkv+ ++ weF+h+sF +g+++l
PDK_30s948451g002 19 FVAKTYQMVNDPRTDAWIRWGNGNNSFLVLDPADFSHLLLPSYFKHSNFSSFVRQLNTYGFKKVDPDR---------WEFAHESFLRGQTHL 101
9****************************************************************999.........*************** PP
XXXXXXXXXX CS
HSF_DNA-bind 94 lekikrkkse 103
l i r+k++
PDK_30s948451g002 102 LPCIVRRKKR 111
******9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 3.9E-37 | 13 | 102 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 6.4E-53 | 15 | 108 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 7.35E-34 | 15 | 108 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| PRINTS | PR00056 | 8.2E-17 | 19 | 42 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Pfam | PF00447 | 5.3E-29 | 19 | 108 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.2E-17 | 57 | 69 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 58 | 82 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 8.2E-17 | 70 | 82 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 332 aa Download sequence Send to blast |
MEEKSSCGKI GYQIQVAPFV AKTYQMVNDP RTDAWIRWGN GNNSFLVLDP ADFSHLLLPS 60 YFKHSNFSSF VRQLNTYGFK KVDPDRWEFA HESFLRGQTH LLPCIVRRKK RNEGGSSSSS 120 SDKKEGIEGE EEEESLFKEL GRLRQEQRAL DEELQRMNKR LQATERRPQQ MMSFLVKVAE 180 DPDFLPRLIL SKKQQLAPEK KRRLVAPSTP PSFSPPPPPP PSLLMASAVS PPLDIDSAAE 240 QTNFSTLEHQ AGPLTLEAER LLLMASSTLV DTGCDGGVGV DNSISYTNVR DSDTVISGNH 300 MRFLPEFGVA ETSSTTQKAV PFPFSLLGHG FV |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2ldu_A | 7e-22 | 16 | 108 | 17 | 120 | Heat shock factor protein 1 |
| 5d5u_B | 7e-22 | 16 | 108 | 26 | 129 | Heat shock factor protein 1 |
| 5d5v_B | 7e-22 | 16 | 108 | 26 | 129 | Heat shock factor protein 1 |
| 5d5v_D | 7e-22 | 16 | 108 | 26 | 129 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA of heat shock promoter elements (HSE). {ECO:0000250}. | |||||
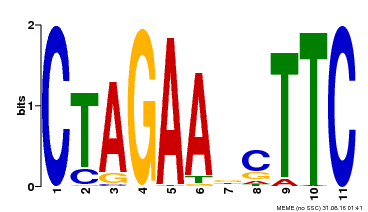
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00379 | DAP | Transfer from AT3G24520 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010924752.2 | 0.0 | heat stress transcription factor C-1b | ||||
| Refseq | XP_029121098.1 | 0.0 | heat stress transcription factor C-1b | ||||
| Swissprot | Q6VBA4 | 1e-79 | HFC1A_ORYSJ; Heat stress transcription factor C-1a | ||||
| TrEMBL | A0A2H3Z8J0 | 1e-155 | A0A2H3Z8J0_PHODC; heat stress transcription factor C-1b-like | ||||
| STRING | XP_008810034.1 | 1e-156 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP11298 | 31 | 41 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24520.1 | 8e-73 | heat shock transcription factor C1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



