 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s931481g009 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | E2F/DP | ||||||||
| Protein Properties | Length: 423aa MW: 46947.4 Da PI: 4.5872 | ||||||||
| Description | E2F/DP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | E2F_TDP | 86.5 | 2e-27 | 98 | 163 | 1 | 71 |
E2F_TDP 1 rkeksLrlltqkflkllekseegivtlnevakeLvsedvknkrRRiYDilNVLealnliekkekneirwkg 71
r+++sL+llt+kf++ll+++++gi++ln++a++L +v ++RRiYDi+NVLe+++liekk kn+irwkg
PDK_30s931481g009 98 RYDSSLGLLTKKFINLLKHAQDGILDLNKAAETL---EV--QKRRIYDITNVLEGIGLIEKKLKNRIRWKG 163
6899******************************...99..****************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 3.5E-29 | 96 | 165 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SuperFamily | SSF46785 | 2.27E-17 | 98 | 161 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM01372 | 1.4E-34 | 98 | 163 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| Pfam | PF02319 | 1.5E-24 | 100 | 163 | IPR003316 | E2F/DP family, winged-helix DNA-binding domain |
| CDD | cd14660 | 9.33E-47 | 174 | 281 | No hit | No description |
| SuperFamily | SSF144074 | 2.12E-32 | 175 | 280 | No hit | No description |
| Pfam | PF16421 | 4.3E-31 | 179 | 278 | IPR032198 | E2F transcription factor, CC-MB domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008284 | Biological Process | positive regulation of cell proliferation | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0010090 | Biological Process | trichome morphogenesis | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0051302 | Biological Process | regulation of cell division | ||||
| GO:0051446 | Biological Process | positive regulation of meiotic cell cycle | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005667 | Cellular Component | transcription factor complex | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 423 aa Download sequence Send to blast |
MPNYLGVPTP EKKIWLFYRL KRKTEQEDNE AAESSEWTTS PGYAEAATSS LLTPVSGKGG 60 RNYGRSKVAK YNKSGPLTPL SNAASPSSNP LTPVGTCRYD SSLGLLTKKF INLLKHAQDG 120 ILDLNKAAET LEVQKRRIYD ITNVLEGIGL IEKKLKNRIR WKGLDDSRPG ELDDDVSSLQ 180 AEVQNLTLQE RSVEDHINEM REKLRVLTED ENNQKWLYVT EADIKGLPCF QNDTLIAIKA 240 PYGTTLEVPD PDEAGEYPHR RYRIVLRSSM GPIDVYLVSQ YEEKFEEMSG VETPQRLPPA 300 SNSGSAENST VEVVTEEGRG KEMAPNVPDC QRMDSDPNAS HDFVGGMMKI VPSDVDTDAD 360 YWLLSDAGVS ITDMWKTAPE VQWDAMGRFN PEDFIASNAS TPRPETPPSG VIDAPSGTNS 420 TQR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1cf7_A | 2e-24 | 93 | 164 | 1 | 74 | PROTEIN (TRANSCRIPTION FACTOR E2F-4) |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds DNA cooperatively with DP proteins through the E2 recognition site, 5'-TTTC[CG]CGC-3' found in the promoter region of a number of genes whose products are involved in cell cycle regulation or in DNA replication. The binding of retinoblastoma-related proteins represses transactivation. Involved in the control of cell-cycle progression from G1 to S phase and from G2 to M phase. Stimulates cell proliferation and delays differentiation. Represses cell enlargement and endoreduplication in auxin-free conditions. {ECO:0000269|PubMed:11669580, ECO:0000269|PubMed:11786543, ECO:0000269|PubMed:11862494, ECO:0000269|PubMed:11891240, ECO:0000269|PubMed:12913157, ECO:0000269|PubMed:16055635, ECO:0000269|PubMed:16514015}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00661 | PBM | Transfer from Pp3c1_6880 | Download |
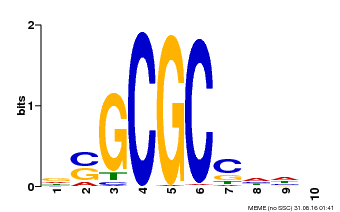
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by light and by auxin. May be up-regulated by E2FA. {ECO:0000269|PubMed:16055635, ECO:0000269|PubMed:16514015, ECO:0000269|PubMed:18424613}. | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HQ413156 | 0.0 | HQ413156.1 Cocos nucifera E2F protein (E2F1) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008784098.1 | 0.0 | transcription factor E2FB-like | ||||
| Swissprot | Q9FV71 | 1e-135 | E2FB_ARATH; Transcription factor E2FB | ||||
| TrEMBL | A0A2H3XI42 | 0.0 | A0A2H3XI42_PHODC; transcription factor E2FB-like | ||||
| STRING | XP_008784098.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP625 | 37 | 148 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G22220.3 | 1e-131 | E2F transcription factor 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



