 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s779741g003 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | GATA | ||||||||
| Protein Properties | Length: 240aa MW: 26843.9 Da PI: 8.8546 | ||||||||
| Description | GATA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | GATA | 55.4 | 8.6e-18 | 168 | 202 | 1 | 35 |
GATA 1 CsnCgttkTplWRrgpdgnktLCnaCGlyyrkkgl 35
C++C + kTp+WR gp g+ktLCnaCG+++++ +l
PDK_30s779741g003 168 CTHCLSHKTPQWRAGPLGPKTLCNACGVRFKSGRL 202
*******************************9986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50114 | 11.121 | 162 | 198 | IPR000679 | Zinc finger, GATA-type |
| SMART | SM00401 | 1.2E-15 | 162 | 216 | IPR000679 | Zinc finger, GATA-type |
| Gene3D | G3DSA:3.30.50.10 | 2.2E-13 | 166 | 200 | IPR013088 | Zinc finger, NHR/GATA-type |
| SuperFamily | SSF57716 | 3.04E-14 | 166 | 226 | No hit | No description |
| CDD | cd00202 | 1.89E-13 | 167 | 224 | No hit | No description |
| PROSITE pattern | PS00344 | 0 | 168 | 193 | IPR000679 | Zinc finger, GATA-type |
| Pfam | PF00320 | 1.5E-15 | 168 | 202 | IPR000679 | Zinc finger, GATA-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007623 | Biological Process | circadian rhythm | ||||
| GO:0009416 | Biological Process | response to light stimulus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 240 aa Download sequence Send to blast |
MEIYCSLSLS LSLSLSLFVG YGFHKKFALQ HLLIEEFEGE EDVSLEWLSI YVEDCLSGTT 60 SYTSQPLPPP TTISHDTPPP KPPTKTSSFR DLAIPAKARS KRRRVTTPKE LTSQDPLLFH 120 QTCWLADSEL ILPKKEQEDK APAVELLVGG GGEAGEKMGK EQQQIRRCTH CLSHKTPQWR 180 AGPLGPKTLC NACGVRFKSG RLLPEYRPAK SPTFVSYKHS NSHKKVMEMR MMAILSSQSN |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 100 | 104 | KRRRV |
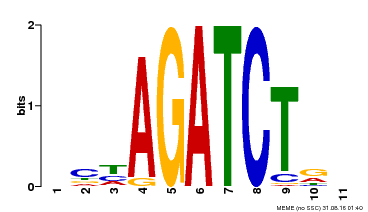
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00048 | PBM | Transfer from AT4G32890 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008812058.1 | 1e-150 | GATA transcription factor 2-like | ||||
| TrEMBL | A0A2H3ZDA6 | 1e-148 | A0A2H3ZDA6_PHODC; GATA transcription factor 2-like | ||||
| STRING | XP_008812058.1 | 1e-149 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5962 | 36 | 56 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G45050.1 | 7e-40 | GATA transcription factor 2 | ||||



