 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s758851g001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 749aa MW: 82482.2 Da PI: 7.1832 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 59.2 | 6.5e-19 | 17 | 75 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+l++ +++ps +r++L +++ +++ +q+kvWFqNrR +ek+
PDK_30s758851g001 17 KYVRYTPEQVEALERLYHDCPKPSSMRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 75
5679*****************************************************97 PP
| |||||||
| 2 | START | 185.7 | 2.4e-58 | 161 | 369 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..gal 91
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v m+++ v+e+l+d++ W ++++++++++v+ g g++
PDK_30s758851g001 161 IAEETLTEFLSKATGTAVEWVQMPGMKPGPDSIGIVAISHGCTGVAARACGLVGMEPT-KVAEILKDRPSWFHDCRSVDVINVLPAGnsGTI 251
789*******************************************************.8888888888*********************** PP
EEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHH CS
START 92 qlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphw 179
+l +++l+a+++l+p Rdf+ +Ry+ l++g++v++++S++s+q p+ +++vRae+lpSg+li+p+++g+s +++v+h+dl+ +++++
PDK_30s758851g001 252 ELLYMQLYAPTTLAPaRDFWLLRYTSVLEDGSLVVCERSLNSTQGGPSmppVQPFVRAEMLPSGYLIRPCEGGGSIMHIVDHMDLEPWSVPE 343
********************************************9998888899************************************** PP
HHHHHHHHHHHHHHHHHHHHTXXXXX CS
START 180 llrslvksglaegaktwvatlqrqce 205
+lr+l++s+++ ++kt++a+l+++++
PDK_30s758851g001 344 VLRPLYESSMVLAQKTTMAALRHLRQ 369
*********************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50071 | 15.58 | 12 | 76 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-16 | 14 | 80 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 4.69E-17 | 15 | 77 | No hit | No description |
| SuperFamily | SSF46689 | 5.13E-17 | 16 | 80 | IPR009057 | Homeodomain-like |
| Pfam | PF00046 | 1.6E-16 | 17 | 75 | IPR001356 | Homeobox domain |
| Gene3D | G3DSA:1.10.10.60 | 1.8E-18 | 19 | 75 | IPR009057 | Homeodomain-like |
| CDD | cd14686 | 9.94E-7 | 69 | 108 | No hit | No description |
| PROSITE profile | PS50848 | 26.098 | 151 | 365 | IPR002913 | START domain |
| CDD | cd08875 | 4.05E-82 | 155 | 370 | No hit | No description |
| Gene3D | G3DSA:3.30.530.20 | 7.1E-25 | 158 | 366 | IPR023393 | START-like domain |
| SuperFamily | SSF55961 | 8.52E-40 | 160 | 371 | No hit | No description |
| SMART | SM00234 | 1.2E-44 | 160 | 370 | IPR002913 | START domain |
| Pfam | PF01852 | 3.8E-56 | 161 | 369 | IPR002913 | START domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 749 aa Download sequence Send to blast |
MAVSSGKEGK TGMDPGKYVR YTPEQVEALE RLYHDCPKPS SMRRQQLIRE CPILSNIEPK 60 QIKVWFQNRR CREKQRKEAS RLQAVNRKLT AMNKLLMEEN DRLQKQVSQL VYENGYFRQQ 120 SRNAALTTPD TSCESVVTSG QHHLTAQRPP RDASPAGLMS IAEETLTEFL SKATGTAVEW 180 VQMPGMKPGP DSIGIVAISH GCTGVAARAC GLVGMEPTKV AEILKDRPSW FHDCRSVDVI 240 NVLPAGNSGT IELLYMQLYA PTTLAPARDF WLLRYTSVLE DGSLVVCERS LNSTQGGPSM 300 PPVQPFVRAE MLPSGYLIRP CEGGGSIMHI VDHMDLEPWS VPEVLRPLYE SSMVLAQKTT 360 MAALRHLRQI VHEVSQSTVT GWGRRPAALR ALSQRLSRGF NEALNGFADD GWSMIGSDGT 420 DDVTILVNSS PSRMMGLNLG FTNGFPSLSN AVLCAKASML LQSSLFLQNV SPAMLLRFLR 480 EHRSEWADSN IDAYSAAAVR ASPCTLPMSR IGCVGGQVIL PLAHTLEHEE FLEVLKLENI 540 GRSQDALMPR DLFLLQLCNG VDENAVGTCS ELVFAPIDAS FADDAPLLPS GFRIIPLDSV 600 MESSSPNRTL DLASTLEIGS TGNRVSGNYS GNSTSTRSVM TIAFQFAFES HLQENVASMA 660 RQYIRNIISS VQRVALALSP SHLGSHGGLR SPPGTPEAIT LARWICYSYR SFSKCMIPFS 720 LVPTDTQYAS SSRTVPHTDT VIEWYWYRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of meristem development to promote lateral organ formation. May regulates procambial and vascular tissue formation or maintenance, and vascular development in inflorescence stems. {ECO:0000269|PubMed:15598805, ECO:0000269|PubMed:15705957, ECO:0000269|PubMed:16617092}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
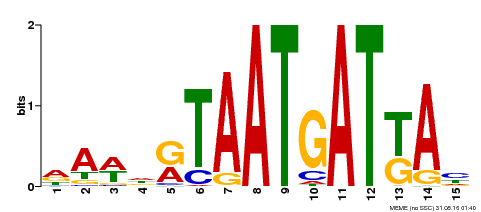
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By auxin. Repressed by miR165 and miR166. {ECO:0000269|PubMed:15773855, ECO:0000269|PubMed:16033795, ECO:0000269|PubMed:16617092, ECO:0000269|PubMed:17237362}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008791633.1 | 0.0 | homeobox-leucine zipper protein ATHB-15-like | ||||
| Swissprot | Q9ZU11 | 0.0 | ATB15_ARATH; Homeobox-leucine zipper protein ATHB-15 | ||||
| TrEMBL | A0A2H3Y0I2 | 0.0 | A0A2H3Y0I2_PHODC; homeobox-leucine zipper protein ATHB-15-like | ||||
| STRING | XP_008791633.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP374 | 38 | 197 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.3 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



