 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s746551g001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 791aa MW: 84897 Da PI: 6.8588 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 66.8 | 2.9e-21 | 119 | 174 | 1 | 56 |
TT--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 1 rrkRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
+++ +++t++q++eLe+lF+++++p++++r eL+++l+L+ rqVk+WFqNrR+++k
PDK_30s746551g001 119 KKRYHRHTPQQIQELEALFKECPHPDEKQRLELSRRLNLESRQVKFWFQNRRTQMK 174
688999***********************************************999 PP
| |||||||
| 2 | START | 183.8 | 9.7e-58 | 327 | 553 | 1 | 206 |
HHHHHHHHHHHHHHHHC-TT-EEEE....EXCCTTEEEEEEESSS......SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-S....EE CS
START 1 elaeeaaqelvkkalaeepgWvkss....esengdevlqkfeeskv.....dsgealrasgvvdmvlallveellddkeqWdetla....ka 79
ela++a++elvk+a+ ep+W s e +n+de+ + f++ + + +ea r++g+v+ ++ lve+l+d + +W +++ +
PDK_30s746551g001 327 ELALAAMDELVKMAQMFEPLWIPSLeggkETLNYDEYERVFPRCIGmkpagFVSEATRETGLVIINSSALVETLMDAA-RWADMFPcmiaRT 417
5899*********************9*9999**********998889*******************************.************* PP
EEEEEECTT......EEEEEEEEXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--..-TTSEE-EESSEEEEEEEECTCEEE CS
START 80 etlevissg......galqlmvaelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe.sssvvRaellpSgiliepksnghsk 163
+t+evissg g+lqlm aelq+lsplvp Rd+ f+R+++q +g w++vdvS+d +++ + + +++++lpSg+++++++ng+sk
PDK_30s746551g001 418 TTTEVISSGmggtrnGTLQLMHAELQVLSPLVPiRDVNFLRFCKQIAEGAWAVVDVSIDGIRQNASaPTPNMKCRRLPSGCVVQDMPNGYSK 509
***********************************************************9999987777889******************** PP
EEEEE-EE--SSXXHHHHHHHHHHHHHHHHHH.HHHHTXXXXXX CS
START 164 vtwvehvdlkgrlphwllrslvksglaegakt.wvatlqrqcek 206
vtwveh++++++ +h+l+r+l++sgl ga++ wvatlqrqce+
PDK_30s746551g001 510 VTWVEHAEYDESAVHQLYRPLLRSGLGLGARRrWVATLQRQCEC 553
******************************988*********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF46689 | 3.89E-21 | 106 | 176 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 3.3E-22 | 109 | 178 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 18.074 | 116 | 176 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 6.4E-19 | 117 | 180 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 2.66E-19 | 119 | 177 | No hit | No description |
| Pfam | PF00046 | 7.5E-19 | 119 | 174 | IPR001356 | Homeobox domain |
| PROSITE pattern | PS00027 | 0 | 151 | 174 | IPR017970 | Homeobox, conserved site |
| PROSITE profile | PS50848 | 37.1 | 318 | 556 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 4.67E-30 | 321 | 553 | No hit | No description |
| CDD | cd08875 | 1.44E-126 | 324 | 552 | No hit | No description |
| Pfam | PF01852 | 2.6E-49 | 327 | 553 | IPR002913 | START domain |
| SMART | SM00234 | 1.7E-41 | 327 | 553 | IPR002913 | START domain |
| SuperFamily | SSF55961 | 4.67E-13 | 581 | 783 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009827 | Biological Process | plant-type cell wall modification | ||||
| GO:0042335 | Biological Process | cuticle development | ||||
| GO:0043481 | Biological Process | anthocyanin accumulation in tissues in response to UV light | ||||
| GO:0048765 | Biological Process | root hair cell differentiation | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 791 aa Download sequence Send to blast |
MSLGGLFDGG SGGARVGAAI PYGSPAMPAG AISQSRLVSP SLPKPLLSSP GLSLALQTNM 60 DGQGDRSLSS IVGGAAGGGG RDADTVGRYK EDENESRSGS DNLDGGSGDD IDQENPRKKK 120 RYHRHTPQQI QELEALFKEC PHPDEKQRLE LSRRLNLESR QVKFWFQNRR TQMKTQMERH 180 ENNILKQEND KLRAENLSIR DAMRNPICSN CGGPVVLGEV SLEEQHLRIE NARLKDELDR 240 VCALAGKFLG KPASALANPL PLPLPSSSLE LAIGTIAFAG LGSAAMTLPV VADFPTAVSS 300 PLGSVGTPVR TVGTLGGVER ERTMLLELAL AAMDELVKMA QMFEPLWIPS LEGGKETLNY 360 DEYERVFPRC IGMKPAGFVS EATRETGLVI INSSALVETL MDAARWADMF PCMIARTTTT 420 EVISSGMGGT RNGTLQLMHA ELQVLSPLVP IRDVNFLRFC KQIAEGAWAV VDVSIDGIRQ 480 NASAPTPNMK CRRLPSGCVV QDMPNGYSKV TWVEHAEYDE SAVHQLYRPL LRSGLGLGAR 540 RRWVATLQRQ CECLAILMAS SLPTRDHNTI TASGRRSMLK LAQRMTDNFC AGVCASSAHK 600 WSKLSVGSIG EDVRVMTRQS VDDPGEPPGV VLSAATSRLR SEWDILSNGG PMQEMAHIAK 660 GQDHGNAVSL LRASAVNANQ SSMLILQETC TDASGSMVVY APVDIPAMHV VMNGGDSAYV 720 ALLPSGFAIL PDGGGGGAQK KVGGSLLTVA FQILVNSQPT AKLTVESVET VNNLISCTVQ 780 KIKAALHCDP Q |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in the regulation of the tissue-specific accumulation of anthocyanins and in cellular organization of the primary root. {ECO:0000269|PubMed:10402424}. | |||||
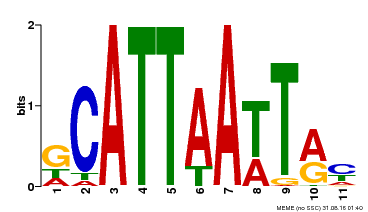
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00421 | DAP | Transfer from AT4G00730 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008799528.1 | 0.0 | homeobox-leucine zipper protein ROC5 | ||||
| Swissprot | Q0WV12 | 0.0 | ANL2_ARATH; Homeobox-leucine zipper protein ANTHOCYANINLESS 2 | ||||
| TrEMBL | A0A2H3YJ62 | 0.0 | A0A2H3YJ62_PHODC; homeobox-leucine zipper protein ROC5 | ||||
| STRING | XP_008799528.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1203 | 37 | 116 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G61150.1 | 0.0 | homeodomain GLABROUS 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



