 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s731461g001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 263aa MW: 29841.7 Da PI: 6.7142 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 53 | 7.6e-17 | 14 | 61 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT+eEd++l ++++G g+W++ ++ g+ R++k+c++rw +yl
PDK_30s731461g001 14 KGPWTPEEDQILTAHIRRYGHGNWRALPKQAGLLRCGKSCRLRWINYL 61
79******************************99************97 PP
| |||||||
| 2 | Myb_DNA-binding | 55.8 | 1.1e-17 | 67 | 112 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
rg++T+eE+e +++++++lG++ W++Ia++++ gRt++++k+ w+++l
PDK_30s731461g001 67 RGNFTKEEEETIIHLHEKLGNR-WSAIAAKLP-GRTDNEIKNVWHTHL 112
89********************.*********.************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-26 | 5 | 64 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 16.676 | 9 | 61 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.22E-31 | 10 | 108 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.6E-14 | 13 | 63 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 5.3E-15 | 14 | 61 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.92E-11 | 16 | 61 | No hit | No description |
| PROSITE profile | PS51294 | 26.542 | 62 | 116 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 5.4E-27 | 65 | 116 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.2E-15 | 66 | 114 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 2.9E-16 | 67 | 112 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.39E-12 | 69 | 112 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0010200 | Biological Process | response to chitin | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 263 aa Download sequence Send to blast |
MGRAPCCEKM GLKKGPWTPE EDQILTAHIR RYGHGNWRAL PKQAGLLRCG KSCRLRWINY 60 LRPDIKRGNF TKEEEETIIH LHEKLGNRWS AIAAKLPGRT DNEIKNVWHT HLKKRVNPNQ 120 VAQESKKKTQ VDAMIEKESK HTPTQLNSGL THPRFDDPRL PPVSPERSYS DISCTATDSS 180 TISRENTCNV SVKEESFSSE EFPVIDESFW SETLSMDFTA LEGPTFGPLG EPFGSFGASN 240 DDDMDFWLRV FMEAGDLQAL PPI |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1h8a_C | 8e-28 | 12 | 116 | 25 | 128 | MYB TRANSFORMING PROTEIN |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator involved in cold stress response (PubMed:14675437, PubMed:20807373, PubMed:22246661). Regulates positively the expression of genes involved in reactive oxygen species (ROS) scavenging such as peroxidase and superoxide dismutase during cold stress. Transactivates a complex gene network that have major effects on stress tolerance and panicle development (PubMed:20807373). {ECO:0000269|PubMed:14675437, ECO:0000269|PubMed:20807373, ECO:0000269|PubMed:22246661}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00375 | DAP | Transfer from AT3G23250 | Download |
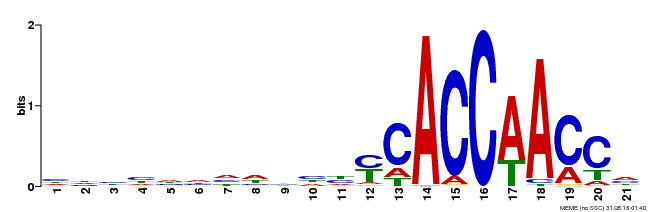
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By cold stress. {ECO:0000269|PubMed:14675437, ECO:0000269|PubMed:20807373, ECO:0000269|Ref.2}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008791593.1 | 0.0 | transcription factor MYB4 | ||||
| Swissprot | Q7XBH4 | 1e-92 | MYB4_ORYSJ; Transcription factor MYB4 | ||||
| TrEMBL | A0A2H3Y121 | 0.0 | A0A2H3Y121_PHODC; transcription factor MYB4 | ||||
| STRING | XP_008791593.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP79 | 38 | 563 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G23250.1 | 2e-81 | myb domain protein 15 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



