 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s6550951g007 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 295aa MW: 33087 Da PI: 9.428 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 36.5 | 1.1e-11 | 24 | 71 | 2 | 47 |
SSS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 2 grWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwqky 47
g+WT+eE +++ da +++ + +W+++a++++ g++ +++s++ ++
PDK_30s6550951g007 24 GAWTQEENKRFEDALARFDRDtpdRWERVAAMIP-GKSVRDIISHYKDL 71
89*****************99*************.***********875 PP
| |||||||
| 2 | Myb_DNA-binding | 47.5 | 4.2e-15 | 130 | 174 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT+eE++l++ + +++G+g+W+ I+r + ++Rt+ q+ s+ qky
PDK_30s6550951g007 130 PWTEEEHKLFLLGLQKYGKGDWRNISRNFVTTRTPTQVASHAQKY 174
8*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 8.907 | 21 | 74 | IPR017884 | SANT domain |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-4 | 22 | 70 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 1.2E-9 | 22 | 74 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 8.97E-12 | 24 | 79 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 8.6E-10 | 24 | 71 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 7.42E-8 | 25 | 71 | No hit | No description |
| PROSITE profile | PS51294 | 17.909 | 123 | 179 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.28E-18 | 125 | 179 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 8.8E-18 | 126 | 177 | IPR006447 | Myb domain, plants |
| Gene3D | G3DSA:1.10.10.60 | 1.1E-12 | 127 | 173 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 5.9E-13 | 127 | 177 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 3.36E-11 | 130 | 175 | No hit | No description |
| Pfam | PF00249 | 9.0E-13 | 130 | 174 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 295 aa Download sequence Send to blast |
MEVLTPRAPY FPNSSWLPGP KRSGAWTQEE NKRFEDALAR FDRDTPDRWE RVAAMIPGKS 60 VRDIISHYKD LEDDVSHIEA GLIPFPGYSS SSFTLDWENS HGYERLKPSY CVGGKRSGGR 120 PDQERKKGVP WTEEEHKLFL LGLQKYGKGD WRNISRNFVT TRTPTQVASH AQKYFIRLNS 180 GGKDKRRSSI HDITTVNLLN NGPPSPSQAS MLAKRSSSAA TTGISDQFSV IVDSNQPNEA 240 AAVFSPSAHG NPFMHPPYEI TSYGMKLPAQ NSQRVILQDS MVADHSMLFQ MHPHG |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 4e-16 | 26 | 91 | 11 | 76 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Involved in the dorsovental asymmetry of flowers. Promotes ventral identity. {ECO:0000269|PubMed:11937495, ECO:0000269|PubMed:9118809}. | |||||
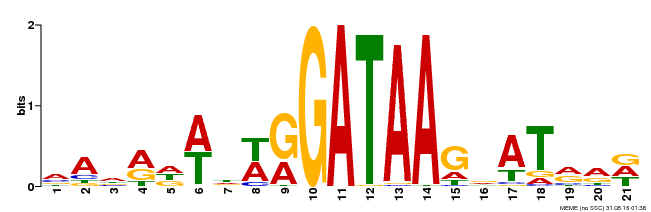
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00296 | DAP | Transfer from AT2G38090 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008785419.1 | 0.0 | transcription factor DIVARICATA isoform X1 | ||||
| Refseq | XP_008785420.1 | 0.0 | transcription factor DIVARICATA isoform X1 | ||||
| Swissprot | Q8S9H7 | 1e-104 | DIV_ANTMA; Transcription factor DIVARICATA | ||||
| TrEMBL | A0A2H3XL35 | 0.0 | A0A2H3XL35_PHODC; transcription factor DIVARICATA isoform X1 | ||||
| STRING | XP_008785419.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2274 | 37 | 96 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G38090.1 | 2e-91 | MYB family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



