 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s6550950g002 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 467aa MW: 52108 Da PI: 5.4167 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 102.8 | 2.1e-32 | 262 | 316 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWt +LHerFveav++L G+ekAtPk +l+lm+v+gLt++hvkSHLQkYRl
PDK_30s6550950g002 262 KPRLRWTLDLHERFVEAVNKLDGAEKATPKGVLKLMNVEGLTIYHVKSHLQKYRL 316
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 9.757 | 259 | 319 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 8.3E-31 | 260 | 317 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 7.35E-17 | 261 | 316 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.1E-24 | 262 | 317 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.8E-8 | 264 | 315 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.7E-20 | 353 | 399 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 467 aa Download sequence Send to blast |
MQIFETPILD ILWKHAGFYG SIEIVFFSQF NCQELLDDKL SCASPSPHVQ RELINSSSPE 60 KGLPFCLKKL CPESEPGSSL SNESHTQHFE HAFSGSSTFC ASLYSSSSTS SESYRQLSNL 120 PFLPHPQKCK LENSVVQSLN SPLLHVGDTS NVHNEDEHSD DLMKDFLNLS GDASDSSFHG 180 ENYDNNSIAL GEQMELQMLS EQLGIAITDN GESPRLDDIY ETLAPSVPLS SNCNQTSQPL 240 TPPAKVQLHS KSSTSTTAAA NKPRLRWTLD LHERFVEAVN KLDGAEKATP KGVLKLMNVE 300 GLTIYHVKSH LQKYRLAKYL PEAQEDKKAS SSEDKKAPTI SHESDLGNKR STQVIEALRM 360 QIEVQKQLHE QLEVQRALQL RIEEHAGYLQ KILEEQQKAS NTFVFSMEMP DTEMQLESPH 420 HSSPHQAESK VDSICPPSSS NSKHKGTDSD TDSKPLEDHK RARLEVE |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 7e-25 | 262 | 319 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 7e-25 | 262 | 319 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 7e-25 | 262 | 319 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 7e-25 | 262 | 319 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 7e-25 | 262 | 319 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 7e-25 | 262 | 319 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 7e-25 | 262 | 319 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 7e-25 | 262 | 319 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in phosphate starvation signaling. Binds to P1BS, an imperfect palindromic sequence 5'-GNATATNC-3', to promote the expression of inorganic phosphate (Pi) starvation-responsive genes. Functionally redundant with PHR1 and PHR2 in regulating Pi starvation response and Pi homeostasis. {ECO:0000250|UniProtKB:Q6YXZ4}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
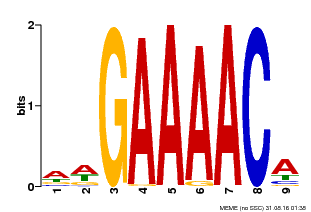
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010927131.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 isoform X1 | ||||
| Refseq | XP_010927132.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 isoform X1 | ||||
| Refseq | XP_010927133.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 isoform X1 | ||||
| Refseq | XP_019707510.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3 isoform X1 | ||||
| Swissprot | A2X0Q0 | 1e-117 | PHR3_ORYSI; Protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| TrEMBL | A0A3Q0I617 | 0.0 | A0A3Q0I617_PHODC; protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| STRING | XP_008796959.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4526 | 37 | 69 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 1e-84 | G2-like family protein | ||||



