 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s6550916g003 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 912aa MW: 104373 Da PI: 6.7383 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 76.9 | 3.9e-24 | 107 | 190 | 2 | 89 |
FAR1 2 fYneYAkevGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHel 89
YneYAk++GF vrk+ ++sk+n+e++ +++vC+kegk++ ++k+ + +++ rtgC+a lkvk++ +++w+v ++ +eHnHel
PDK_30s6550916g003 107 LYNEYAKHMGFGVRKKLCRRSKTNNEMIAAWYVCTKEGKSAANEKA----ECSQSARRTGCRAALKVKRMANKEWKVVHFAKEHNHEL 190
7****************************************99888....889999*******************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 1.0E-20 | 107 | 190 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 3.3E-30 | 303 | 396 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 10.311 | 594 | 630 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 1.2E-7 | 604 | 629 | IPR007527 | Zinc finger, SWIM-type |
| SMART | SM00575 | 1.0E-7 | 605 | 632 | IPR006564 | Zinc finger, PMZ-type |
| PRINTS | PR00600 | 1.1E-6 | 766 | 793 | IPR000009 | Protein phosphatase 2A regulatory subunit PR55 |
| PRINTS | PR00600 | 1.1E-6 | 794 | 821 | IPR000009 | Protein phosphatase 2A regulatory subunit PR55 |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000159 | Cellular Component | protein phosphatase type 2A complex | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| GO:0008601 | Molecular Function | protein phosphatase type 2A regulator activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 912 aa Download sequence Send to blast |
MESTNDGSSQ KQANVEDIEL AAEIGLVDDD DDSDDDDDHD DDDDDEEEEE EEEAASLIRG 60 SAAIELADPS TDLEVSSLAG NPDEVGGTRS EPSIGMVFQS AVAAYRLYNE YAKHMGFGVR 120 KKLCRRSKTN NEMIAAWYVC TKEGKSAANE KAECSQSARR TGCRAALKVK RMANKEWKVV 180 HFAKEHNHEL DLENVCLFRS HMKNRISNRS RINMAGVGHK YAMAAPGKQS GGQDGECQNH 240 MDRGQRRFFA PADAHAIHML FIHMHSKNPA FFYAVEFDGE QHLKNVLWAD AKAKVAYTHF 300 GYVVTFDTTY LTDGYKVPFA PFVGVNHHGQ SILLGCALVA DQSTASFVWL FKRWLDAMSG 360 RPPKAIISDY NKDIAAAISQ VFPETRHRYC LWQILKRVPE KLGDVCEARE NFLKKLNKCI 420 YDSTTADEFE RRWWKLIHRF GLVNEEWLQS LYKDRQQWVP LYLKDTFFAG MSISQRNESV 480 STIFHKFITR ETGLKELLDK YEVALESKFE DEAQEDFWSF HSKTKERAHP SPFELQLADV 540 YTRNMFEKFQ MEVLGISSVF ASKSEENGDI ATYVVKAYEV QEKRKTRRLL EKEYKVVWNT 600 MERKVCCACR LFEFKGYLCR HALAVFLALG VLEVPSFYIL KRWTKDAKSR HVLDEGYTVH 660 GDCSESVAQR FNDICVRSFK FAEEASVTKR SYDVALLSLR EAFRRVTHAN NSVRKSCQRN 720 YPHNGVSNIA NEPSAKFQPR AENGTSKQNN NVSRSYEQLG HNLVMFREVP KGLRHLLISQ 780 TFISAENLRV NLWNLEISNQ YFNIDDMRPL DMEDLVVVRD AKGRGAVAVS HMLIVLQGTH 840 VKTMDDRVAG LQMSKHSHIA CAWSMKLQTL LLHKYFVGQA WRLGDELQDV QQVYKNLMAE 900 LEAVDEVKVE NS |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 581 | 588 | EKRKTRRL |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
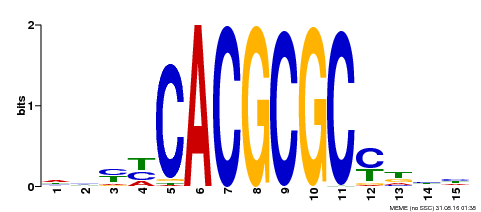
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_026662207.1 | 0.0 | protein FAR-RED IMPAIRED RESPONSE 1-like isoform X1 | ||||
| TrEMBL | A0A3Q0IA20 | 0.0 | A0A3Q0IA20_PHODC; protein FAR-RED IMPAIRED RESPONSE 1-like isoform X1 | ||||
| STRING | XP_008796002.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP18 | 32 | 967 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 1e-166 | FAR1 family protein | ||||



