 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s1094441g001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 364aa MW: 39586.3 Da PI: 7.9687 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 99.6 | 2.1e-31 | 194 | 249 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+p+LH++F++a++qLGGs+ AtPk+i+elmkv++Lt+++vkSHLQkYRl+
PDK_30s1094441g001 194 KARRCWSPDLHRQFIHALQQLGGSHAATPKQIRELMKVDDLTNDEVKSHLQKYRLH 249
68****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.873 | 191 | 251 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-27 | 192 | 252 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.97E-16 | 192 | 251 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 5.3E-26 | 194 | 249 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.8E-6 | 196 | 247 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 364 aa Download sequence Send to blast |
MDLAERARRC HEYVEALEEE RRKIQVFPRE LPLCLQLVTE GSYLSLLFVR AFCGAFILGS 60 IRVQMGSSEE TVTDGPVLEE FIPLKPTSSS IEEEKGGSME MDQERSDNKP DWLRSVQLWS 120 QDPDHSLPKA EPPKKPIAVN AKKIGGAFRP FDREKHVPAP VPSAAAPASS TTVGGGSSGE 180 KEKDKEGQSQ PHRKARRCWS PDLHRQFIHA LQQLGGSHAA TPKQIRELMK VDDLTNDEVK 240 SHLQKYRLHS RRPSPAVQSS GTSSPPTPQY VVVGEIWVRP PDYAAAAVAT AAQPANGAGG 300 IYAPVASLPS DLRHQQQQVK QQSHRSPTGP LHSQGRCSGD DGAANSASPT TSSSSQTTTA 360 SPPF |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 1e-12 | 194 | 252 | 2 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 1e-12 | 194 | 252 | 2 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 1e-12 | 194 | 247 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 1e-12 | 194 | 247 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 1e-12 | 194 | 247 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 1e-12 | 194 | 247 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 1e-12 | 194 | 252 | 2 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 1e-12 | 194 | 252 | 2 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 1e-12 | 194 | 252 | 2 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 1e-12 | 194 | 252 | 2 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 1e-12 | 194 | 252 | 2 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 1e-12 | 194 | 252 | 2 | 60 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional repressor that may play a role in response to nitrogen. May be involved in a time-dependent signaling for transcriptional regulation of nitrate-responsive genes. Binds specifically to the DNA sequence motif 5'-GAATC-3' or 5'-GAATATTC-3'. Represses the activity of its own promoter trough binding to these motifs. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
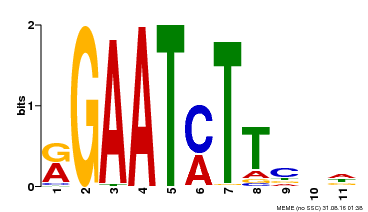
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by nitrate. {ECO:0000269|PubMed:23324170}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008799311.1 | 0.0 | transcription factor NIGT1-like isoform X1 | ||||
| Swissprot | Q6Z869 | 6e-88 | NIGT1_ORYSJ; Transcription factor NIGT1 | ||||
| TrEMBL | A0A2H3YJ60 | 0.0 | A0A2H3YJ60_PHODC; transcription factor NIGT1-like isoform X1 | ||||
| STRING | XP_008799311.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP6470 | 37 | 54 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 8e-75 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



