 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | PDK_30s1055981g001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Arecales; Arecaceae; Coryphoideae; Phoeniceae; Phoenix
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 866aa MW: 96119 Da PI: 6.9688 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 71.1 | 1.4e-22 | 159 | 260 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkl 84
f+k+lt sd++++g +++ +++a+e+ +++++ +l +d++g +W++++i+r++++r++l++GW+ Fv++++L +gD ++F
PDK_30s1055981g001 159 FCKTLTASDTSTHGGFSVLRRHADEClppldmcRQPPTQ--ELAAKDLHGVEWRFRHIFRGQPRRHLLQSGWSVFVSSKRLVAGDAFIFL- 246
99*****************************87555544..8************************************************. PP
-SSSEE..EEEEE-S CS
B3 85 dgrsefelvvkvfrk 99
+ +++el+v+v+r+
PDK_30s1055981g001 247 -RGENGELRVGVRRA 260
.449999*****996 PP
| |||||||
| 2 | Auxin_resp | 120.6 | 1.2e-39 | 285 | 367 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha++t+++F+v+Y+Pr+s+seF+v++++++++lk++ svGmRfkm+fe+e+++e+r+ Gt+vg++d+d+ rWp+SkWr+Lk
PDK_30s1055981g001 285 AWHAVNTGTMFTVYYKPRTSPSEFIVSYDQYMESLKNNHSVGMRFKMRFEGEEAPEQRFIGTIVGIEDADSNRWPGSKWRCLK 367
89*******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 6.1E-41 | 152 | 274 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 6.67E-40 | 153 | 288 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.56E-20 | 157 | 259 | No hit | No description |
| PROSITE profile | PS50863 | 11.462 | 159 | 261 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 3.8E-22 | 159 | 261 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.2E-20 | 159 | 260 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.2E-36 | 285 | 367 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 2.0E-10 | 728 | 824 | IPR033389 | AUX/IAA domain |
| PROSITE profile | PS51745 | 24.11 | 739 | 823 | IPR000270 | PB1 domain |
| SuperFamily | SSF54277 | 5.89E-12 | 741 | 816 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 866 aa Download sequence Send to blast |
MASSEVSVVK GSNGRGESFS SGCSGPNQGA SVAMETGPGF AAAPAAGKES EDALYTELWH 60 ACAGPLVTVP RVGEKVFYFP QGHMEQVEAS TNQVADQQMP IYNLPSKILC RVINVQLKAE 120 PDTDEVFAQV TLLPEKQDDN AVEKETLPPP PPRPRVHSFC KTLTASDTST HGGFSVLRRH 180 ADECLPPLDM CRQPPTQELA AKDLHGVEWR FRHIFRGQPR RHLLQSGWSV FVSSKRLVAG 240 DAFIFLRGEN GELRVGVRRA MRQQTNVPSS VISSHSMHLG VLATAWHAVN TGTMFTVYYK 300 PRTSPSEFIV SYDQYMESLK NNHSVGMRFK MRFEGEEAPE QRFIGTIVGI EDADSNRWPG 360 SKWRCLKVRW DETSSISRPE RVSPWKIEPA LTPPPINPLP MPRSKRPRSN VVPCSPDSSV 420 LTKEAASKIN IDPSHSHGTP RVLQGQEVMT LRSSFAENSK SDAAQKPIMW SSSCDEEKTD 480 ASVQRRLGSE KWMQMVQHER NYMDMLSGFW PTHDSHGFNS TLFEQNPGDA HFLRSYFQDP 540 EGSRNALPGH WSLMPPNLSY TSNEPSLKMT AQPGEQSYQK AGNIRYVGQV GYSALPGLGI 600 EQHSPNCLAH FLSTSLMENP SQPRVVKPQP SGAAENDVVK PQVSGNCKLF GFHLNSNPTA 660 SETVTLQQMN ASTHEPEPHT HAAASLNQLQ AKEANRYSEP FKVIKSADQT PVGSELEKSF 720 QPCPQTSKDV QSKHSGSARS CTKVHKQGIA LGRSVDLTKF NGYDELFAEL DRKFEFQGEL 780 ISPNKNWLVV YTDNEGDMML VGDDPWHEFC SMVRKIFIYT REEVQRMNPG TLTSRVEESP 840 ASSEEITAAK ERKGQVPASS AHPESC |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-179 | 52 | 389 | 17 | 354 | Auxin response factor 1 |
| 4ldw_A | 1e-179 | 52 | 389 | 17 | 354 | Auxin response factor 1 |
| 4ldw_B | 1e-179 | 52 | 389 | 17 | 354 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
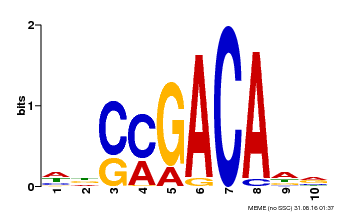
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_008811924.1 | 0.0 | auxin response factor 2A-like | ||||
| Refseq | XP_026655713.1 | 0.0 | auxin response factor 2A-like | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A2H3ZCV6 | 0.0 | A0A2H3ZCV6_PHODC; Auxin response factor | ||||
| STRING | XP_008811924.1 | 0.0 | (Phoenix dactylifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1178 | 36 | 111 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



