 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr040390.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 456aa MW: 49531.8 Da PI: 9.8865 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 45.6 | 1.5e-14 | 353 | 402 | 5 | 54 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkk 54
+r+rr++kNRe+A rsR+RK+a++ eLe +v++L++ N++L+k+ ee+ +
Pbr040390.1 353 RRQRRMIKNRESAARSRARKQAYTLELEAEVAKLKEMNEELQKKQEEIME 402
79****************************************99888765 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 2.3E-14 | 349 | 417 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.955 | 351 | 402 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.5E-10 | 353 | 401 | No hit | No description |
| CDD | cd14707 | 1.73E-26 | 353 | 407 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 2.0E-14 | 353 | 402 | No hit | No description |
| Pfam | PF00170 | 6.1E-13 | 353 | 402 | IPR004827 | Basic-leucine zipper domain |
| PROSITE pattern | PS00036 | 0 | 356 | 371 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 456 aa Download sequence Send to blast |
MGSNFNFKSF GEAAPVEGNG GRAGGSFGLV RQSSVYSLTF DEFQNTIGGL GKDYGSMNMD 60 ELLKNIWTAE ETQGMTATSG VGGEVNAPGS NLQRQGSLTL PRTLSQKTVD EVWKNLIRET 120 TDYKYNNVAA GSNLPQRQQT LGEMTLEDFL LKAGVVREDI QPPVVRPNNG GFYGELYRPK 180 NNVGLASGFQ QPNISNGILG NRVADNKNSV PNQSPTLALN VGGVRTSQQQ TLQLPPQQQP 240 LFPKPSTMAF APSMHLVNNA QLSSPRTRGP MAGVVEPSMN TAFSQAGGFP GAGIGTVGLG 300 TGGGAVATRS PANQISPDVI AKSSGDTSSL SPVPYMFNRG RKCSGAVEKV VERRQRRMIK 360 NRESAARSRA RKQAYTLELE AEVAKLKEMN EELQKKQEEI MEVQKDQVHH YVGDNEAAMG 420 RQKAMLTTNI DRPLVTRMKL DCEYKMSARV YERYR* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Binds to the ABA-responsive element (ABRE). Mediates stress-responsive ABA signaling. {ECO:0000269|PubMed:11884679, ECO:0000269|PubMed:15361142}. | |||||
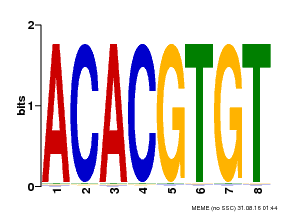
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00038 | PBM | Transfer from AT4G34000 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr040390.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by drought, salt, abscisic acid (ABA). {ECO:0000269|PubMed:10636868, ECO:0000269|PubMed:16284313}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009346257.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| Refseq | XP_009346258.1 | 0.0 | PREDICTED: ABSCISIC ACID-INSENSITIVE 5-like protein 5 | ||||
| Swissprot | Q9M7Q3 | 1e-103 | AI5L6_ARATH; ABSCISIC ACID-INSENSITIVE 5-like protein 6 | ||||
| TrEMBL | A0A498K3J6 | 0.0 | A0A498K3J6_MALDO; Uncharacterized protein | ||||
| STRING | XP_009346256.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1962 | 34 | 81 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G34000.2 | 1e-94 | abscisic acid responsive elements-binding factor 3 | ||||



