 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr037761.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 973aa MW: 107916 Da PI: 6.479 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 131.4 | 3.2e-41 | 143 | 220 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqve+C+adls+ak+yhrrhkvC++hska+++lv +++qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+++
Pbr037761.1 143 VCQVEDCKADLSSAKDYHRRHKVCDMHSKATKALVGNVMQRFCQQCSRFHALQEFDEGKRSCRRRLAGHNRRRRKTNP 220
6**************************************************************************976 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 4.3E-33 | 138 | 205 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 32.129 | 141 | 218 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 3.27E-38 | 142 | 223 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 1.9E-29 | 144 | 217 | IPR004333 | Transcription factor, SBP-box |
| Gene3D | G3DSA:1.25.40.20 | 7.0E-6 | 259 | 273 | IPR020683 | Ankyrin repeat-containing domain |
| SuperFamily | SSF48403 | 1.23E-6 | 753 | 855 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.62E-5 | 789 | 854 | No hit | No description |
| Gene3D | G3DSA:1.25.40.20 | 7.0E-6 | 790 | 855 | IPR020683 | Ankyrin repeat-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 973 aa Download sequence Send to blast |
MESEFGGMGG VMVPDLTGVG KRSMEWDLNG WKWDGDLFTA SPLNSTPSDG RSRQFFPVRP 60 ETLFDAGLSN SSSGSDNIGP GNEKGTRELE KRRRDVFVEN GELNDEAASL NLKLGGQTYP 120 VMEEEVQTGK KTKIIGTTSN RAVCQVEDCK ADLSSAKDYH RRHKVCDMHS KATKALVGNV 180 MQRFCQQCSR FHALQEFDEG KRSCRRRLAG HNRRRRKTNP DTAVNGGSLN NESGSSYLLI 240 SLLRILSNMH SSSSDQTKDQ DVVSHLLRSL ANVAGMADGR NVSKLLQGSQ GLFNSGTSVQ 300 TARKVLDMDD GVNPEDPLRP KGHCSILPAS RDSSESKSVT PEPTSRRFQL NDIDLNSTYD 360 DSQDFVENLG NSHVPASPGT ASPGFPSWMQ RDSHKSSPPQ TSGNSDLTST QSPSSSSGEA 420 QSRTDRIVFK LFGKDPSELP FALRSQILDW LSHSPTNIES YIRPGCIILT IYLRLEKSTW 480 EEFCCHLGSS LKTLLDAADD PFWRTGWVYT RVQHFVAFTY NGEVVLDTPL PLKTDKSCRI 540 SCIKPIAISL SERAEFVVKG LNLSRSTTRL LCALEGKYLV QETCYDLMDD ADSSVEDDEQ 600 QCLRFSCSIP NITGRGFIEV EDHGLSSSFF PFIVAEQEVC SEICMLEDVI EAAETDDDIQ 660 SGPEKVEAKN QALDFIHELG WLLHRSRVKF RLGQLDPNLD LFPFGRFRLL MEFSIDHDWC 720 AVVKKLLGIL FDGTVDAGKH PSVESALLDM HLLHRAVRRN GRHMASPPQS GSEQNKQVDK 780 DGNSFLFKPD VFGPMGLTPL HIAASTDGCE QVLDALTDDP GKVGIKAWKN ARDSTGLTPY 840 DYACLRSHYS YVQIVQRKIS KTLESGHVVL DIPGLTLDRN GKQKQSDADK SSRVASLETE 900 KNEIKAILRH CRLCEQKPAH STTRSLVYRP AMLSMVIVAA VCVCVALLFK STPEVLFVFE 960 PFRWEHLKFG SS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 9e-30 | 143 | 217 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00633 | PBM | Transfer from PK06791.1 | Download |
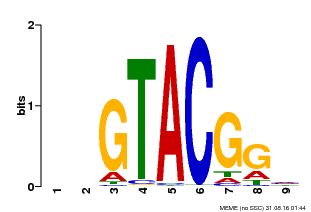
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr037761.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009359778.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A498HJS4 | 0.0 | A0A498HJS4_MALDO; Uncharacterized protein | ||||
| STRING | XP_009359778.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF1539 | 34 | 95 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



