 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr029607.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 557aa MW: 61013 Da PI: 6.9503 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 51.8 | 1.4e-16 | 353 | 399 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCCC...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPkaskapskKlsKaeiLekAveYIksLq 55
hn Er+RRdriN+++ L+el+P++ K +Ka++L +A+eY+ksLq
Pbr029607.1 353 VHNLSERKRRDRINEKMKALQELIPHS-----NKTDKASMLDEAIEYLKSLQ 399
6*************************7.....5******************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF47459 | 2.49E-20 | 347 | 413 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| PROSITE profile | PS50888 | 18.12 | 349 | 398 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.07E-17 | 352 | 403 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 3.7E-20 | 353 | 407 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 5.6E-14 | 353 | 399 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 3.5E-17 | 355 | 404 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009704 | Biological Process | de-etiolation | ||||
| GO:0010161 | Biological Process | red light signaling pathway | ||||
| GO:0010244 | Biological Process | response to low fluence blue light stimulus by blue low-fluence system | ||||
| GO:0010600 | Biological Process | regulation of auxin biosynthetic process | ||||
| GO:0010928 | Biological Process | regulation of auxin mediated signaling pathway | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 557 aa Download sequence Send to blast |
MNSCIPEWNF ESDLPLTNQK KSIGPDHELV ELLWRNGQVV LNSQTHRKPS FNPNDQSRQV 60 QKHDQQSIRS EGLYGNSSNP IQDDDTVSLI HYPLEDSFDK EFCSHFFSEF PSCDPIELDK 120 PIKQFEGEKV VKFDASDTAR LVSNSPRPNG NSSAGVEYAA NPMPPPRYQY INTDQQNQNQ 180 GGVGKIVNFS QFATPGRVSA GSSRGQLGGK ESGNLAQAEV RECSVMTVGS SYVGSNQVLN 240 DLDVSRASSN CDGTTGLSAG HFYDNVHKIM PQSETGKTDT LDPTLTSSSG GSGSSFGRGG 300 NQSNVVNSNK RKGREAADSE CQSEAAELES ASAARSKSAH RSGSTRRSRA AEVHNLSERK 360 RRDRINEKMK ALQELIPHSN KTDKASMLDE AIEYLKSLQM QLQVMWMGSG MAPMMFPGVQ 420 HYISRMGMGM GMGMGPPALP SMQNPMHLPR HPLVDQCMNV APAANQTVMC QPPVLNPIDY 480 HNQMQNPSFQ EQYARLMNFH HMQTMSQPMN MFRFGSQPLQ QNKMMAPNGF SSGPFGGGAA 540 TNDPLSGKMS KSLEST* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 357 | 362 | ERKRRD |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00606 | ChIP-seq | Transfer from AT2G43010 | Download |
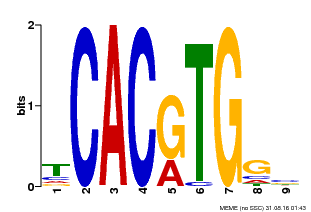
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr029607.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KP689415 | 0.0 | KP689415.1 Prunus pseudocerasus isolate Unigene5738 bHLH transcription factor mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009356576.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| Refseq | XP_009356577.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| Refseq | XP_009356578.1 | 0.0 | PREDICTED: transcription factor PIF4-like | ||||
| TrEMBL | A0A498JA55 | 0.0 | A0A498JA55_MALDO; Uncharacterized protein | ||||
| STRING | XP_009356576.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8146 | 29 | 44 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G59060.4 | 6e-40 | phytochrome interacting factor 3-like 6 | ||||



