 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr027474.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 377aa MW: 41592 Da PI: 7.2656 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.1 | 7e-32 | 211 | 266 | 1 | 56 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRla 56
k+r++W+peLH+rF++a++qLGGs+ AtPk+i+elmkv+gLt+++vkSHLQkYRl+
Pbr027474.1 211 KQRRNWSPELHRRFLNALQQLGGSHAATPKQIRELMKVDGLTNDEVKSHLQKYRLH 266
79****************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.384 | 208 | 268 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-28 | 209 | 269 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 5.18E-18 | 209 | 269 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.3E-27 | 211 | 266 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 7.2E-8 | 213 | 264 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 377 aa Download sequence Send to blast |
MESTEEMQAS SKLGFREYVK ALEEEMHKIQ VFKRELPLCY ELVTHAIERC KQQVSDTTAD 60 YMHGQSECSQ QTSSEGHVFE EFIPLKRTSS SDSEDDEVHE SQEAKDNEKD DKIAGGDKKK 120 SDWLRSVQLW NTTPDAPLKE ELPRKALVME VNRNGGAFQP FQKEKSIGKT NGPVAKQPSS 180 AAATSSTTDT VSGSSGENNK KEEKDGQGQR KQRRNWSPEL HRRFLNALQQ LGGSHAATPK 240 QIRELMKVDG LTNDEVKSHL QKYRLHTRRP TPTMRSNNNA AHAPQFVVVG GIWVPPQDYA 300 AVAATTASGE AANNGIYAPV ASTPAVTQVS PPVQRLRLKQ PEPSHSDERV SHSEGRGHSN 360 STATSSSTHT HISPAF* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 2e-14 | 211 | 264 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 2e-14 | 211 | 264 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_A | 2e-14 | 211 | 264 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 2e-14 | 211 | 264 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 2e-14 | 211 | 264 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 2e-14 | 211 | 264 | 1 | 54 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 2e-14 | 211 | 264 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 2e-14 | 211 | 264 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 2e-14 | 211 | 264 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 2e-14 | 211 | 264 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 2e-14 | 211 | 264 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 2e-14 | 211 | 264 | 2 | 55 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor involved in phosphate signaling in roots. {ECO:0000250|UniProtKB:Q9FX67}. | |||||
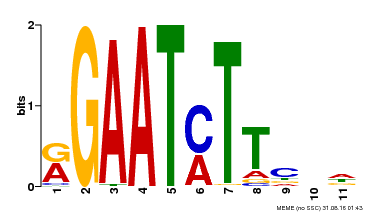
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00160 | DAP | Transfer from AT1G25550 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr027474.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009336613.1 | 0.0 | PREDICTED: myb family transcription factor EFM | ||||
| Swissprot | Q9FPE8 | 1e-103 | HHO3_ARATH; Transcription factor HHO3 | ||||
| TrEMBL | A0A498I6H4 | 0.0 | A0A498I6H4_MALDO; Uncharacterized protein | ||||
| STRING | XP_009336613.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF3918 | 33 | 63 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G25550.1 | 3e-89 | G2-like family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



