 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr016948.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | HSF | ||||||||
| Protein Properties | Length: 346aa MW: 38770.4 Da PI: 9.2483 | ||||||||
| Description | HSF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HSF_DNA-bind | 109.5 | 2.5e-34 | 13 | 104 | 2 | 102 |
HHHHHHHHHCTGGGTTTSEESSSSSEEEES-HHHHHHHTHHHHSTT--HHHHHHHHHHTTEEE---SSBTTTTXTTSEEEEESXXXXXXXXXXXXXX CS
HSF_DNA-bind 2 FlkklyeiledeelkeliswsengnsfvvldeeefakkvLpkyFkhsnfaSFvRQLnmYgFkkvkdeekkskskekiweFkhksFkkgkkellekik 98
F+ k+y++++d+++++li+w + +nsf+v+d+ f++++Lp yFkh+nf+SFvRQLn+YgF+kv+ ++ weF++++F +g+++ll+++
Pbr016948.1 13 FVLKTYQMVNDPTTDNLITWGRANNSFIVVDPLVFSQRLLPAYFKHNNFSSFVRQLNTYGFRKVDPDK---------WEFANEWFLRGQTHLLRNVA 100
9********************999*****************************************999.........******************** PP
XXXX CS
HSF_DNA-bind 99 rkks 102
r+k+
Pbr016948.1 101 RRKQ 104
*997 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.10 | 2.5E-36 | 5 | 96 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| SMART | SM00415 | 3.2E-52 | 9 | 102 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| SuperFamily | SSF46785 | 9.66E-33 | 10 | 102 | IPR011991 | Winged helix-turn-helix DNA-binding domain |
| Pfam | PF00447 | 5.8E-29 | 13 | 102 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.5E-18 | 13 | 36 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.5E-18 | 51 | 63 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PROSITE pattern | PS00434 | 0 | 52 | 76 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| PRINTS | PR00056 | 2.5E-18 | 64 | 76 | IPR000232 | Heat shock factor (HSF)-type, DNA-binding |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 346 aa Download sequence Send to blast |
MMMEESNNVI APFVLKTYQM VNDPTTDNLI TWGRANNSFI VVDPLVFSQR LLPAYFKHNN 60 FSSFVRQLNT YGFRKVDPDK WEFANEWFLR GQTHLLRNVA RRKQMGKSSN SNSLFSNSNS 120 NFLPTKHEEL NDEDIIMEIS RLKQEQKALE EEIGNMNRRL ETTERRPQQM MAFLHKVAED 180 PEILPRMMLE KDRATTALLG EKKRRGVTRM MLEKDRATTA LLGEKKRRVM MIASTSSSSS 240 GMGATTSVRT EDEDDGTVGV ISSSPEAGFE MESFNRYSTS PEVQTASDWL RQRRFVDQVR 300 AVQNPNNMNP TASGHGAENI RNSSSSGYGY GNGNGNGGGP GGGGG* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5hdg_A | 2e-24 | 10 | 102 | 7 | 110 | Heat shock factor protein 1 |
| 5hdn_A | 2e-24 | 10 | 102 | 7 | 110 | Heat shock factor protein 1 |
| 5hdn_B | 2e-24 | 10 | 102 | 7 | 110 | Heat shock factor protein 1 |
| 5hdn_C | 2e-24 | 10 | 102 | 7 | 110 | Heat shock factor protein 1 |
| 5hdn_D | 2e-24 | 10 | 102 | 7 | 110 | Heat shock factor protein 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional regulator that specifically binds DNA sequence 5'-AGAAnnTTCT-3' known as heat shock promoter elements (HSE). | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00379 | DAP | Transfer from AT3G24520 | Download |
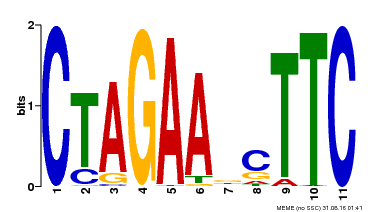
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr016948.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_018506836.1 | 0.0 | PREDICTED: heat stress transcription factor C-1 isoform X1 | ||||
| Swissprot | Q9LV52 | 7e-86 | HSFC1_ARATH; Heat stress transcription factor C-1 | ||||
| TrEMBL | A0A498HV72 | 0.0 | A0A498HV72_MALDO; Uncharacterized protein | ||||
| STRING | XP_009374015.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF8367 | 30 | 38 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G24520.1 | 4e-86 | heat shock transcription factor C1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



