 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr005887.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | SRS | ||||||||
| Protein Properties | Length: 358aa MW: 38264.9 Da PI: 7.9576 | ||||||||
| Description | SRS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF702 | 221.3 | 1.9e-68 | 122 | 269 | 3 | 153 |
DUF702 3 sgtasCqdCGnqakkdCaheRCRtCCksrgfdCathvkstWvpaakrrerqqqlaaasskaaasaaeaaskrkrelkskkqsalsstklssaeskkel 100
g+++CqdCGnqakkdC h RCRtCCksrgf+C+thvkstWvpaakrrerq+q +a+++++++++ ++ + r ++ k ++ ++l++ +++s +s l
Pbr005887.1 122 GGGINCQDCGNQAKKDCPHLRCRTCCKSRGFQCQTHVKSTWVPAAKRRERQEQFSALQQNRNQQQ-QQIQFRGENPKRQRDNSLACVRIPSDTSG--L 216
5789**************************************************99988887655.44555555556889999*******99866..8 PP
DUF702 101 etsslPeevsseavfrcvrvssvddgeeelaYqtavsigGhvfkGiLydqGle 153
e + +P+evss+avfrcv+vs++dd +e++aYqtav+igGhvfkG+LydqG++
Pbr005887.1 217 ELNPFPPEVSSPAVFRCVKVSAMDDVDEQFAYQTAVNIGGHVFKGLLYDQGID 269
8899***********************************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05142 | 8.8E-66 | 124 | 269 | IPR007818 | Protein of unknown function DUF702 |
| TIGRFAMs | TIGR01623 | 1.3E-26 | 126 | 168 | IPR006510 | Zinc finger, lateral root primordium type 1 |
| TIGRFAMs | TIGR01624 | 9.2E-26 | 221 | 268 | IPR006511 | Lateral Root Primordium type 1, C-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009299 | Biological Process | mRNA transcription | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 358 aa Download sequence Send to blast |
MAGFFCLGGR DRSPNSRGKQ EGENQSGRVG GVAAEAAAGG GGGNLFLYRS TGEAQAEIYN 60 KGFEIWPSQY HPHQNLNYYS FGVDPSYNRR HLDNNNDGVS ADDLSAGLRL TVMRGGGLGS 120 SGGGINCQDC GNQAKKDCPH LRCRTCCKSR GFQCQTHVKS TWVPAAKRRE RQEQFSALQQ 180 NRNQQQQQIQ FRGENPKRQR DNSLACVRIP SDTSGLELNP FPPEVSSPAV FRCVKVSAMD 240 DVDEQFAYQT AVNIGGHVFK GLLYDQGIDG QYYPSSGGGG ESSSGGQGTS HDHHHDGGGS 300 GGGRGHPHHH QQHNLVTVAA AGGTSGNPST TLLDPSLFPA PLNAFMAGTQ YFPPPRS* |
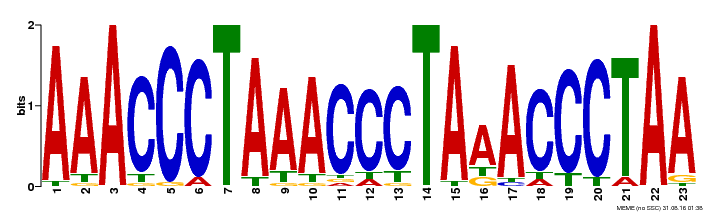
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00152 | DAP | Transfer from AT1G19790 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr005887.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009335611.1 | 0.0 | PREDICTED: protein SHI RELATED SEQUENCE 1 isoform X1 | ||||
| TrEMBL | A0A498HUT3 | 0.0 | A0A498HUT3_MALDO; Uncharacterized protein | ||||
| STRING | XP_009335611.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF882 | 34 | 125 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G75520.1 | 6e-72 | SHI-related sequence 5 | ||||



