 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Pbr003518.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; rosids; fabids; Rosales; Rosaceae; Maloideae; Maleae; Pyrus
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 514aa MW: 56421 Da PI: 6.4603 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 32.6 | 1.7e-10 | 209 | 250 | 4 | 45 |
XCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 4 lkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaL 45
k rr+++NReAAr+sR+RKka++++Le +L++ ++L
Pbr003518.1 209 AKTLRRLAQNREAARKSRLRKKAYVQQLESSRIKLTQLEQDL 250
6889*************************9777777666665 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.20.5.170 | 1.7E-7 | 205 | 252 | No hit | No description |
| SMART | SM00338 | 7.7E-7 | 206 | 269 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 9.186 | 208 | 252 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 9.0E-8 | 209 | 251 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 1.0E-6 | 210 | 252 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 213 | 228 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF14144 | 8.5E-32 | 291 | 366 | IPR025422 | Transcription factor TGA like domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 514 aa Download sequence Send to blast |
MAASHRVGDA TAATPTATTA GLSSELGPSN HHVPYAVLHG MNASSTSFIN QEGSAFDFGE 60 LEEAIARQVR NDEAQAPLFT GGGGAGRPAA TLEMFPSWPM RFHQTPGGSS KSGEEESTDS 120 GSQVNTLTTT TTTTTNQLEL EPESPMSKNP SSSQAAFDQK HLQFQQQQQQ QEMAISDTGP 180 NSQTPSAQKP SQEKRKGAGS TSENKQLDAK TLRRLAQNRE AARKSRLRKK AYVQQLESSR 240 IKLTQLEQDL QRARAQGLFL GGCGGGLGNI SSGAAIFDME YARWLEDDHR HMSELRTGLQ 300 AHLSDSDLRL VVDGYVSHYD EIFQLKGMAA KSDVFHLITG MWTTPAERCF LWMGGFRPSE 360 LIKMLIAQLD PLTEQQFMGI YNLQQSSQQG EEALTQGLEQ LHQSLVDTIA GGPVIDGMQQ 420 MAVALGKLTN LEGFVRQADN LRQQTLHQLR RILTVRQAAR CFLVIGEYYG RLRALSSLWA 480 SRPRETMISD DNSCQTTTDL QMLHPSQNHF ASF* |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Together with TGA10, basic leucine-zipper transcription factor required for anther development, probably via the activation of SPL expression in anthers and via the regulation of genes with functions in early and middle tapetal development (PubMed:20805327). Required for signaling responses to pathogen-associated molecular patterns (PAMPs) such as flg22 that involves chloroplastic reactive oxygen species (ROS) production and subsequent expression of H(2)O(2)-responsive genes (PubMed:27717447). {ECO:0000269|PubMed:20805327, ECO:0000269|PubMed:27717447}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00131 | DAP | Transfer from AT1G08320 | Download |
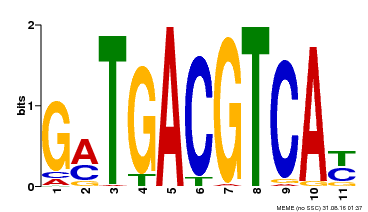
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Pbr003518.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by flg22 in leaves. {ECO:0000269|PubMed:27717447}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | KC770002 | 1e-179 | KC770002.1 Pyrus pyrifolia microsatellite TXY004 sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009364312.1 | 0.0 | PREDICTED: transcription factor TGA2.2-like | ||||
| Swissprot | Q93XM6 | 0.0 | TGA9_ARATH; Transcription factor TGA9 | ||||
| TrEMBL | A0A498KBI7 | 0.0 | A0A498KBI7_MALDO; Uncharacterized protein | ||||
| STRING | XP_009364312.1 | 0.0 | (Pyrus x bretschneideri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Fabids | OGEF4555 | 33 | 57 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G08320.3 | 0.0 | bZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



