 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peaxi162Scf00058g02017.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia
|
||||||||
| Family | SBP | ||||||||
| Protein Properties | Length: 969aa MW: 108142 Da PI: 6.0645 | ||||||||
| Description | SBP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SBP | 130.5 | 6e-41 | 107 | 184 | 1 | 78 |
--SSTT-----TT--HHHHHTT--HHHHT-S-EEETTEEEEE-TTTSSEEETTT--SS--S-STTTT-------S--- CS
SBP 1 lCqvegCeadlseakeyhrrhkvCevhskapvvlvsgleqrfCqqCsrfhelsefDeekrsCrrrLakhnerrrkkqa 78
+Cqv++C +dls+ k+yhrrhkvCe+hska+++lv++++qrfCqqCsrfh+l+efDe+krsCrrrLa+hn+rrrk+qa
Peaxi162Scf00058g02017.1 107 VCQVDDCGVDLSKGKDYHRRHKVCEMHSKATTALVANVMQRFCQQCSRFHALEEFDEGKRSCRRRLAGHNKRRRKTQA 184
6**************************************************************************986 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:4.10.1100.10 | 3.1E-31 | 104 | 169 | IPR004333 | Transcription factor, SBP-box |
| PROSITE profile | PS51141 | 31.787 | 105 | 182 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF103612 | 1.44E-37 | 106 | 186 | IPR004333 | Transcription factor, SBP-box |
| Pfam | PF03110 | 2.7E-30 | 108 | 181 | IPR004333 | Transcription factor, SBP-box |
| SuperFamily | SSF48403 | 7.19E-7 | 743 | 856 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 9.8E-5 | 745 | 857 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.11E-5 | 746 | 856 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 969 aa Download sequence Send to blast |
MEGETQLYYR PMDLREVTNK NIEWDPNDWK WDAHLFIAKP NNYQQTANLA LVSSNTNTSS 60 SCSDEVNLEK RRRVLIDENG GNLSLKLFDG NTNNGKRSKV TSCNRAVCQV DDCGVDLSKG 120 KDYHRRHKVC EMHSKATTAL VANVMQRFCQ QCSRFHALEE FDEGKRSCRR RLAGHNKRRR 180 KTQADSGDNN NKTLSDGQAS GYSLMSLLKI LSNTHSNGTN HLEDQDLLSH LVRSLASQGA 240 LNGDNNLSGL LQATSKLLNN DMSPSRNSDI VSLVSNGSRG PPRPKEQHFA TLAAQMPQRS 300 SDVHHVRLDD VVTVSCQSPG TPFPIQNSSP AYAQDSESPA GRSKVINFDL NDVYVDSDDG 360 GENLERSPLP ADRGTGSLEC RSWLQQDSLQ SSPPQTSRNS DSASAHSPSS SSSDAQSRTD 420 RIVFKLFGKD PSDFPFVLRA QILDWLSHSP TEIESYIRPG CVVLTVYLRL PESAWEELFC 480 DLSSSLTRLL DVRGDSFWRK GWIYVMLQNQ IALVHDGQLL LDVPLPIETI EHTSIISVKP 540 IAVPASDRAQ FIVRGFNLTK PSTRLLCALE GDYLVAETND EAEHVDDIDK HEKLQSINLT 600 CSIPAISGRG FIEVEDHGLS NGFFPFIIAE DDVCSEIRML ESVIELTSAD YINGLTNNAD 660 LKSQAMDFIY EMGWLLHRNN VKARLGNSGP HAVLFPLKRF KWLMEFSVDR EWCAVVRKLL 720 NLLLDGTVGT EENSFLKVSL SEMGLLHKAV RRNSRPLVEL LLTYTPEGIA DELCSEYQSL 780 VGINGNFLFR PDNVGPAGLT PLHIAAGIDG CEDVLDALTD DPGMVALEAW KNTRDSTGFT 840 PEDYARLRGH YSYIHLVQRK INRKASSRHI VVDIPSILSD ENSNQKDEYA VASLEISETD 900 RRPIRPCGLC DRKLAYGSRS RTLLYRPAMF SMVALAAVCV CVALLFKSSP EVLYIFRPFR 960 WEMLDFGSS |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ul4_A | 2e-30 | 107 | 181 | 10 | 84 | squamosa promoter binding protein-like 4 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 69 | 73 | KRRRV |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Trans-acting factor that binds specifically to the consensus nucleotide sequence 5'-TNCGTACAA-3' of AP1 promoter. Binds specifically to the 5'-GTAC-3' core sequence. {ECO:0000269|PubMed:10524240, ECO:0000269|PubMed:16095614}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00603 | PBM | Transfer from AT2G47070 | Download |
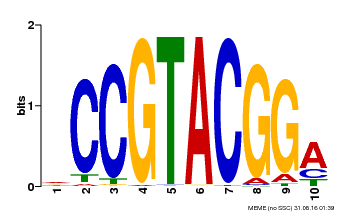
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_016507693.1 | 0.0 | PREDICTED: squamosa promoter-binding-like protein 1 | ||||
| Swissprot | Q9SMX9 | 0.0 | SPL1_ARATH; Squamosa promoter-binding-like protein 1 | ||||
| TrEMBL | A0A2K9ZXI5 | 0.0 | A0A2K9ZXI5_PETHY; Squamosa promoter binding-like protein 12c | ||||
| STRING | XP_009773067.1 | 0.0 | (Nicotiana sylvestris) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA1871 | 24 | 66 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G47070.1 | 0.0 | squamosa promoter binding protein-like 1 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



