 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Peaxi162Scf00001g00012.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Solanales; Solanaceae; Petunioideae; Petunia
|
||||||||
| Family | CAMTA | ||||||||
| Protein Properties | Length: 1074aa MW: 120744 Da PI: 5.5653 | ||||||||
| Description | CAMTA family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | CG-1 | 172.5 | 5.8e-54 | 22 | 147 | 3 | 118 |
CG-1 3 ke.kkrwlkneeiaaiLenfekheltlelktrpksgsliLynrkkvryfrkDGyswkkkkdgktvrEdhekLKvgg......... 77
+e ++rwl++ ei++iL+n++++++t e++ rp sgs++L++rk++ryfrkDG++w+kkkdgktv+E+hekLKvg
Peaxi162Scf00001g00012.1 22 SEaQHRWLRPAEICEILQNYRNFHITPEAPYRPVSGSVFLFDRKVLRYFRKDGHNWRKKKDGKTVKEAHEKLKVGVdslgigvgs 106
4449**********************************************************************9999******* PP
CG-1 78 vevlycyYahseenptfqrrcywlLeeelekivlvhylevk 118
++vl+cyYah+ee+ +fqrr+yw+Le++l +iv+vhylevk
Peaxi162Scf00001g00012.1 107 IDVLHCYYAHGEEDDNFQRRSYWMLEQDLMHIVFVHYLEVK 147
**************************************985 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51437 | 77.13 | 17 | 152 | IPR005559 | CG-1 DNA-binding domain |
| SMART | SM01076 | 2.2E-76 | 20 | 147 | IPR005559 | CG-1 DNA-binding domain |
| Pfam | PF03859 | 5.2E-46 | 23 | 146 | IPR005559 | CG-1 DNA-binding domain |
| Gene3D | G3DSA:2.60.40.10 | 4.8E-6 | 490 | 577 | IPR013783 | Immunoglobulin-like fold |
| SuperFamily | SSF81296 | 7.14E-18 | 491 | 577 | IPR014756 | Immunoglobulin E-set |
| Pfam | PF01833 | 9.6E-8 | 491 | 576 | IPR002909 | IPT domain |
| SuperFamily | SSF48403 | 1.87E-16 | 680 | 782 | IPR020683 | Ankyrin repeat-containing domain |
| Gene3D | G3DSA:1.25.40.20 | 1.8E-16 | 683 | 784 | IPR020683 | Ankyrin repeat-containing domain |
| CDD | cd00204 | 1.35E-14 | 686 | 780 | No hit | No description |
| PROSITE profile | PS50088 | 8.79 | 688 | 720 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50297 | 17.422 | 688 | 782 | IPR020683 | Ankyrin repeat-containing domain |
| SMART | SM00248 | 4300 | 688 | 717 | IPR002110 | Ankyrin repeat |
| PROSITE profile | PS50088 | 9.484 | 721 | 753 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 0.0036 | 721 | 750 | IPR002110 | Ankyrin repeat |
| SMART | SM00248 | 3300 | 760 | 789 | IPR002110 | Ankyrin repeat |
| SuperFamily | SSF52540 | 6.64E-7 | 891 | 946 | IPR027417 | P-loop containing nucleoside triphosphate hydrolase |
| SMART | SM00015 | 7.3 | 895 | 917 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 7.895 | 896 | 925 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 0.0041 | 897 | 916 | IPR000048 | IQ motif, EF-hand binding site |
| SMART | SM00015 | 0.0016 | 918 | 940 | IPR000048 | IQ motif, EF-hand binding site |
| PROSITE profile | PS50096 | 9.432 | 919 | 943 | IPR000048 | IQ motif, EF-hand binding site |
| Pfam | PF00612 | 2.0E-4 | 921 | 940 | IPR000048 | IQ motif, EF-hand binding site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009409 | Biological Process | response to cold | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0071275 | Biological Process | cellular response to aluminum ion | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 1074 aa Download sequence Send to blast |
MIMAGSGSDP PGFRLDIKQI LSEAQHRWLR PAEICEILQN YRNFHITPEA PYRPVSGSVF 60 LFDRKVLRYF RKDGHNWRKK KDGKTVKEAH EKLKVGVDSL GIGVGSIDVL HCYYAHGEED 120 DNFQRRSYWM LEQDLMHIVF VHYLEVKVRL YNFFPYQGNK ANVSFNRSIE SAHSNYVKEY 180 SLSDTFHGSH KKLASLKADS ASLASTVTSA YEEAESEDSH QVCSRFHSYP EQVSGMDSRL 240 VENRDTISKS YGSPLSSVEY PSLPGIDGGG KCDLGNSASG PQRTNDLGSR APVFERCSNG 300 EVVSTHDFKN NMPVHGNWQY SLEGSPLQFL GQSVNQDLIA DSCYDLLNGF HSENLSSNQY 360 TVRGQSYLYP DEQDEQQTEL NFQNLNSPME VQADIIQENF MDMLGLGDYS TIKQPRLGSV 420 KLEEGLNKSD SFSRWVAKEL EDVEELHMQS TDRLSWNVID IEDDCSCLPS QLHVDSDSLV 480 PSLSQEQLFS IIDFSPNWAY SSLETKVLIT GRFLKSDCKV VECKWSCMFG EVEVPAEVLA 540 DGVLRCHAPP HKPGVLPFYV TCSNRLACSE VREFEYRLGP SREFGASNVS AIEMHLLERF 600 ESVMSLEPVS SCYSSDSPEA APEKQSTVNK IICMMEEDMA DRPSDFDTSQ SKVKEDLFLE 660 RKLRQNFYVW LVHQVTADGK GRTVLDDEGQ GVLHLAAALG YDWALRPILA SGVSVDFRDM 720 NGWTALHWAA FYGREKTVVG LVSLGASPGA LTDPSAEFPL GRTPADLASA NRHKGISGFL 780 AESSLTSHLS KLTVTDAKEE LASEVLGAKV GETVTERVAV TSTGDDVPDV LSLKDSLAAI 840 RNATQAAARI HQIFRVQSFQ RKQIIENNDN ELSSDEHALS VVASRTCKLA QNNGIAHAAA 900 IQIQKKFRGW NRRKEFLLIR QRIVKIQAHV RGHQVRKKYK PIIWSVGILE KVILRWRRKG 960 SGLRGFRSEA VMNKPSTEDD SLPEDDYDFL KEGRKQTEVR MQKALARVKS MTQYPEARAQ 1020 YRRLLTAAEG LRETKEDGPT QVLENQKDAS YAEEDLFDVD NLLDEDTFMS IAFD |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cxk_A | 1e-13 | 491 | 581 | 9 | 93 | calmodulin binding transcription activator 1 |
| 2cxk_B | 1e-13 | 491 | 581 | 9 | 93 | calmodulin binding transcription activator 1 |
| 2cxk_C | 1e-13 | 491 | 581 | 9 | 93 | calmodulin binding transcription activator 1 |
| 2cxk_D | 1e-13 | 491 | 581 | 9 | 93 | calmodulin binding transcription activator 1 |
| 2cxk_E | 1e-13 | 491 | 581 | 9 | 93 | calmodulin binding transcription activator 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the DNA consensus sequence 5'-[ACG]CGCG[GTC]-3' (By similarity). Regulates transcriptional activity in response to calcium signals (Probable). Binds calmodulin in a calcium-dependent manner (By similarity). Involved in freezing tolerance in association with CAMTA1 and CAMTA3. Contributes together with CAMTA1 and CAMTA3 to the positive regulation of the cold-induced expression of DREB1A/CBF3, DREB1B/CBF1 and DREB1C/CBF2 (PubMed:23581962). Involved together with CAMTA3 and CAMTA4 in the positive regulation of a general stress response (PubMed:25039701). Involved in tolerance to aluminum. Binds to the promoter of ALMT1 transporter and contributes to the positive regulation of aluminum-induced expression of ALMT1 (PubMed:25627216). {ECO:0000250|UniProtKB:Q8GSA7, ECO:0000269|PubMed:23581962, ECO:0000269|PubMed:25039701, ECO:0000269|PubMed:25627216, ECO:0000305|PubMed:11925432}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00043 | PBM | Transfer from AT5G64220 | Download |
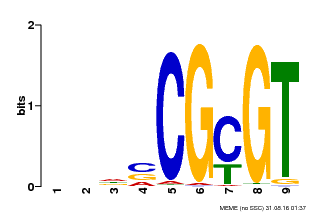
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By salt, wounding, abscisic acid, H(2)O(2) and salicylic acid (PubMed:12218065). Induced by aluminum (PubMed:25627216). {ECO:0000269|PubMed:12218065, ECO:0000269|PubMed:25627216}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_009600617.1 | 0.0 | PREDICTED: calmodulin-binding transcription activator 2-like isoform X1 | ||||
| Swissprot | Q6NPP4 | 0.0 | CMTA2_ARATH; Calmodulin-binding transcription activator 2 | ||||
| TrEMBL | A0A1S4BAF3 | 0.0 | A0A1S4BAF3_TOBAC; calmodulin-binding transcription activator 2-like isoform X1 | ||||
| STRING | XP_009600617.1 | 0.0 | (Nicotiana tomentosiformis) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA2230 | 21 | 46 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64220.2 | 0.0 | Calmodulin-binding transcription activator protein with CG-1 and Ankyrin domains | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



