 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MA_2237g0020 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Picea
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 789aa MW: 85857.2 Da PI: 7.1619 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 73.3 | 3e-23 | 138 | 239 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.......---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSSEE CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.......ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrsef 90
f k+lt+sd+++ g +++p+ +ae++ + ++ + t+ +d +g++W++++iyr++++r++lt+GW+ Fv++++L +gD++vF ++ ++
MA_2237g0020 138 FAKTLTQSDANNGGGFSVPRYCAETIfpkldysSDPPVQ--TVLAKDVHGEVWKFRHIYRGTPRRHLLTTGWSTFVNQKKLVAGDSIVFL--RSLNG 230
89****************************995444444..8999*********************************************..44789 PP
..EEEEE-S CS
B3 91 elvvkvfrk 99
el+v+++r+
MA_2237g0020 231 ELCVGIRRS 239
99****997 PP
| |||||||
| 2 | Auxin_resp | 111.3 | 9.5e-37 | 317 | 399 | 2 | 83 |
Auxin_resp 2 ahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrl.sGtvvgvsdldpvrWpnSkWrsLk 83
a+ a+t+++FevvY+Prast+eF+vk++ v +al++k+++GmRfkmafetedss++++ +Gt++ v+++dp++Wp S+Wr+L+
MA_2237g0020 317 ATLAATGQPFEVVYYPRASTPEFCVKANAVDTALRMKWAAGMRFKMAFETEDSSRISWfMGTISLVQCADPIHWPTSPWRLLQ 399
57899****************************************************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:2.40.330.10 | 1.2E-36 | 129 | 245 | IPR015300 | DNA-binding pseudobarrel domain |
| SuperFamily | SSF101936 | 4.71E-29 | 133 | 248 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 1.99E-21 | 137 | 238 | No hit | No description |
| SMART | SM01019 | 4.1E-22 | 138 | 240 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.816 | 138 | 240 | IPR003340 | B3 DNA binding domain |
| Pfam | PF02362 | 2.4E-21 | 138 | 239 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 8.5E-32 | 317 | 399 | IPR010525 | Auxin response factor |
| PROSITE profile | PS51745 | 17.315 | 703 | 783 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0007389 | Biological Process | pattern specification process | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0048829 | Biological Process | root cap development | ||||
| GO:0051301 | Biological Process | cell division | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 789 aa Download sequence Send to blast |
MPVPCTMNGP RITAMDSAHG YGLVMQRGAD DGSPGLDSQL WHACAGGMVQ MPSVNARVYY 60 FPQGQAEHSA MNVEFPASLR AKPAILCRVL NVKFMADTET DEVYARIRLV PIKPNEAAME 120 DCSEEIDNAA ANEKPVSFAK TLTQSDANNG GGFSVPRYCA ETIFPKLDYS SDPPVQTVLA 180 KDVHGEVWKF RHIYRGTPRR HLLTTGWSTF VNQKKLVAGD SIVFLRSLNG ELCVGIRRST 240 RSTGGGDSSA WHLAGQQQQQ GSCMSPYRSS SRWEVKGGDN FSAAFFSGGD KASANGNSNG 300 FARNKGKVST KSVIESATLA ATGQPFEVVY YPRASTPEFC VKANAVDTAL RMKWAAGMRF 360 KMAFETEDSS RISWFMGTIS LVQCADPIHW PTSPWRLLQV TWDEPDLLQN VKRVSPWLVE 420 VVSNVPPIQL SPFALPKKKL RVSQHPEFQF EGHGFMGGLQ MATLTNNVLG QINPWHGLSE 480 NVPAGMQGAR HGHIYGISLS DFHPNKVQSG LSLDSFHQLE QGAPLSTPVS TELNIGCFSQ 540 LERSSVQDNL SCVLTMGNSS QSEQKATKGR SSSSTTKSAS FLLFGKPIHT EQSPKSEQQQ 600 QSGVSSSDGP GLHAVNDSGS PGLTSNSSTD GNPEVVDRLQ RAMTGRSDTS KLSENYGLRL 660 YCGETLDSTV NNGGGNLSWF KDRQVSLPTL ENGSGGKTPE DGILHCKAFL ESEDVGRTLD 720 LSVFSSYEQL HNRLAKMFGI EDSDLSNRVL YKDAAGVGRH TGDQPYGDFM KTVRRLTILS 780 DSSSDNMGR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 4e-72 | 33 | 420 | 16 | 353 | Auxin response factor 1 |
| 4ldw_A | 4e-72 | 33 | 420 | 16 | 353 | Auxin response factor 1 |
| 4ldw_B | 4e-72 | 33 | 420 | 16 | 353 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). | |||||
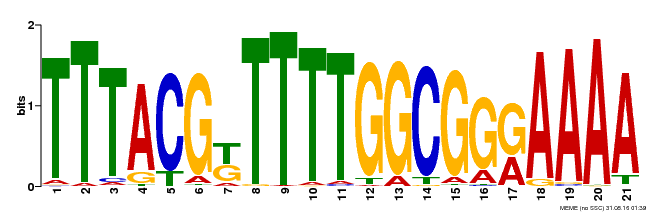
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00461 | DAP | Transfer from AT4G30080 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT108545 | 0.0 | BT108545.1 Picea glauca clone GQ03123_F06 mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010253698.1 | 0.0 | PREDICTED: auxin response factor 18-like | ||||
| Swissprot | Q653H7 | 0.0 | ARFR_ORYSJ; Auxin response factor 18 | ||||
| TrEMBL | A0A2P1NR35 | 0.0 | A0A2P1NR35_9SPER; Auxin response factor | ||||
| STRING | XP_010253698.1 | 0.0 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP526 | 15 | 78 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G30080.1 | 0.0 | auxin response factor 16 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



