 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MA_19520g0020 | ||||||||
| Common Name | HB-3 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Picea
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 822aa MW: 89658 Da PI: 6.1092 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 57.1 | 3e-18 | 18 | 76 | 3 | 57 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHC....TS-HHHHHHHHHHHHHHHHC CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkkl....gLterqVkvWFqNrRakekk 57
k ++t+eq+e+Le+++ ++++ps +r++L +++ +++ +q+kvWFqNrR +ek+
MA_19520g0020 18 KYVRYTSEQVEALERVYSECPKPSSLRRQQLIRECpilsNIEPKQIKVWFQNRRCREKQ 76
6789*****************************************************97 PP
| |||||||
| 2 | START | 163.8 | 1.2e-51 | 144 | 352 | 2 | 205 |
HHHHHHHHHHHHHHHC-TT-EEEEEXCCTTEEEEEEESSS.SCEEEEEEEECCSCHHHHHHHHHCCCGGCT-TT-SEEEEEEEECTT..EEEEEEE CS
START 2 laeeaaqelvkkalaeepgWvkssesengdevlqkfeeskvdsgealrasgvvdmvlallveellddkeqWdetlakaetlevissg..galqlmv 95
+aee+++e+++ka+ ++ Wv+++ +++g++++ +++ s++++g a+ra+g+v +++ v+e+l+d++ W ++++ ++l + +g g+++l +
MA_19520g0020 144 IAEETLAEFLSKAKGAAVDWVQMPGMKPGPDSIGIVAISNTCNGVAARACGLVGLDPT-KVAEILKDRPSWLRDCRCLDVLTAFPTGngGTIELLY 238
7899******************************************************.8888888888*****************999******* PP
EXXTTXX-SSX.EEEEEEEEEEE.TTS-EEEEEEEEE-TTS--....-TTSEE-EESSEEEEEEEECTCEEEEEEEE-EE--SSXXHHHHHHHHHH CS
START 96 aelqalsplvp.RdfvfvRyirqlgagdwvivdvSvdseqkppe...sssvvRaellpSgiliepksnghskvtwvehvdlkgrlphwllrslvks 187
++++a ++l++ Rdf+++Ry+ l++g++v++++S++ +q p+ +++vRae+lpSg+li+p+++g+s + +v+h+dl+ ++++++lr+l++s
MA_19520g0020 239 MQTYAATTLASaRDFWTLRYTTVLEDGSLVVCERSLSGTQGGPSippVQHFVRAEMLPSGYLIQPCEGGGSIIRIVDHMDLEPWSVPEVLRPLYES 334
********9999*****************************99999999*********************************************** PP
HHHHHHHHHHHHTXXXXX CS
START 188 glaegaktwvatlqrqce 205
+++ ++k++ a+l+r+++
MA_19520g0020 335 STVLAQKMTIAALRRLRQ 352
*************99876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.6E-18 | 6 | 76 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 15.775 | 13 | 77 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.1E-14 | 15 | 81 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 5.13E-16 | 18 | 80 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 7.92E-16 | 18 | 78 | No hit | No description |
| Pfam | PF00046 | 7.4E-16 | 18 | 76 | IPR001356 | Homeobox domain |
| PROSITE profile | PS50848 | 25.731 | 134 | 349 | IPR002913 | START domain |
| CDD | cd08875 | 3.02E-74 | 138 | 354 | No hit | No description |
| SuperFamily | SSF55961 | 4.94E-36 | 143 | 355 | No hit | No description |
| SMART | SM00234 | 2.1E-37 | 143 | 353 | IPR002913 | START domain |
| Gene3D | G3DSA:3.30.530.20 | 1.6E-23 | 144 | 349 | IPR023393 | START-like domain |
| Pfam | PF01852 | 3.7E-49 | 144 | 352 | IPR002913 | START domain |
| Pfam | PF08670 | 4.4E-47 | 679 | 821 | IPR013978 | MEKHLA |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009965 | Biological Process | leaf morphogenesis | ||||
| GO:0010014 | Biological Process | meristem initiation | ||||
| GO:0010075 | Biological Process | regulation of meristem growth | ||||
| GO:0010087 | Biological Process | phloem or xylem histogenesis | ||||
| GO:0048263 | Biological Process | determination of dorsal identity | ||||
| GO:0080060 | Biological Process | integument development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0008289 | Molecular Function | lipid binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 822 aa Download sequence Send to blast |
MAVMANNKDA KNSMDTSKYV RYTSEQVEAL ERVYSECPKP SSLRRQQLIR ECPILSNIEP 60 KQIKVWFQNR RCREKQRKEA SRLQTKQVSQ LVYENGYMRQ QLQNASVAAT DTSCESVVTS 120 GQHQHNPTPQ HPPRDASPAG LLSIAEETLA EFLSKAKGAA VDWVQMPGMK PGPDSIGIVA 180 ISNTCNGVAA RACGLVGLDP TKVAEILKDR PSWLRDCRCL DVLTAFPTGN GGTIELLYMQ 240 TYAATTLASA RDFWTLRYTT VLEDGSLVVC ERSLSGTQGG PSIPPVQHFV RAEMLPSGYL 300 IQPCEGGGSI IRIVDHMDLE PWSVPEVLRP LYESSTVLAQ KMTIAALRRL RQIAQEATGE 360 VVFGWGRQPA VLRTFSQRLS RGFNEAVNGF TDDGWSLLGS DGVEDVTIAI NSSPNKHFAS 420 QVNASNGLTT LGGGILCAKA SMLLQNVPPA LLVRFLREHR SEWADSNIDA YSAAALKASP 480 YSVPGSRAGG FSGSQVILPL AHTVEHEEFL EVIKLEGHGL TQEEAVLSRD MFLLQLCSGI 540 DENAAGACAE LVFAPIDESF ADDAPLLPSG FRVIPLESRT DGSGGPNRTL DLASALEVGS 600 AGTRTSGDSG ANSNLRSVLT IAFQFTYESH LRENVAAMAR QYVRSVVASV QRVAMALAPS 660 RLSAHVGPRP PPGTPEALTL ARWICQSYRL HIGVDLFRAD CEASESVLKL LWHHSDAIMC 720 CSVKSLPVFT FANQAGLDML ETTLVALQDI SLDKILDENG RKSFFTDYGQ IIQQGYAYLP 780 AGICLSSMGR PASYDRAIAW KVLNDEDSTH CIVFMFMNWS FM |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
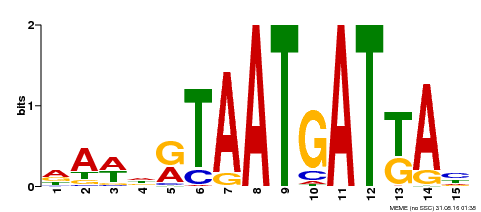
| Motif ID | Method | Source | Motif file |
| MP00200 | DAP | Transfer from AT1G52150 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | HQ391914 | 0.0 | HQ391914.1 Picea glauca class III homeodomain leucine zipper protein (HB3) mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010247499.1 | 0.0 | PREDICTED: homeobox-leucine zipper protein HOX32-like | ||||
| Swissprot | Q6AST1 | 0.0 | HOX32_ORYSJ; Homeobox-leucine zipper protein HOX32 | ||||
| TrEMBL | Q0QSS2 | 0.0 | Q0QSS2_PICGL; Homeodomain-leucine zipper trancription factor HB-3 | ||||
| TrEMBL | Q0QUA8 | 0.0 | Q0QUA8_PICMA; Homeodomain-leucine zipper trancription factor HB-3 | ||||
| TrEMBL | Q0QUG0 | 0.0 | Q0QUG0_PICAB; Homeodomain-leucine zipper trancription factor HB-3 | ||||
| STRING | XP_010247498.1 | 0.0 | (Nelumbo nucifera) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP651 | 16 | 71 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G52150.2 | 0.0 | HD-ZIP family protein | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



