 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MA_10432580g0010 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Picea
|
||||||||
| Family | ARF | ||||||||
| Protein Properties | Length: 873aa MW: 96862.6 Da PI: 6.0445 | ||||||||
| Description | ARF family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | B3 | 75 | 8.9e-24 | 161 | 262 | 1 | 99 |
EEEE-..-HHHHTT-EE--HHH.HTT.....---..--SEEEEEETTS-EEEEEE..EEETTEEEE-TTHHHHHHHHT--TT-EEEEEE-SSS CS
B3 1 ffkvltpsdvlksgrlvlpkkfaeeh.....ggkkeesktltledesgrsWevkliyrkksgryvltkGWkeFvkangLkegDfvvFkldgrs 88
f+k+lt sd++++g +++ +++a+e+ +++++ s++l+ +d++g +W++++i+r++++r++lt+GW+ Fv++++L +gD ++F + +
MA_10432580g0010 161 FCKTLTASDTSTHGGFSVLRRHADEClppldMSQQPPSQELVAKDLHGVEWRFRHIFRGQPRRHLLTTGWSVFVSSKRLVAGDAFIFL--RGE 251
99*****************************55566678*************************************************..449 PP
EE..EEEEE-S CS
B3 89 efelvvkvfrk 99
++el+v+v+r+
MA_10432580g0010 252 NGELRVGVRRA 262
999*****996 PP
| |||||||
| 2 | Auxin_resp | 123.1 | 1.9e-40 | 287 | 369 | 1 | 83 |
Auxin_resp 1 aahaastksvFevvYnPrastseFvvkvekvekalkvkvsvGmRfkmafetedsserrlsGtvvgvsdldpvrWpnSkWrsLk 83
a+ha+st+++F+v+Y+Pr+s+seF++++++++ka+k+++svGmRfkm+fe+e+++e+r++Gt+vg+ d+dpv+W++SkWr+Lk
MA_10432580g0010 287 ASHAVSTRTMFSVYYKPRTSPSEFIIPYDQYMKAMKINFSVGMRFKMKFEGEEVPEQRFTGTIVGICDVDPVSWRDSKWRCLK 369
789******************************************************************************96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101936 | 1.31E-43 | 154 | 290 | IPR015300 | DNA-binding pseudobarrel domain |
| Gene3D | G3DSA:2.40.330.10 | 2.7E-42 | 155 | 276 | IPR015300 | DNA-binding pseudobarrel domain |
| CDD | cd10017 | 6.44E-22 | 160 | 261 | No hit | No description |
| Pfam | PF02362 | 1.2E-21 | 161 | 262 | IPR003340 | B3 DNA binding domain |
| PROSITE profile | PS50863 | 12.012 | 161 | 263 | IPR003340 | B3 DNA binding domain |
| SMART | SM01019 | 8.4E-24 | 161 | 263 | IPR003340 | B3 DNA binding domain |
| Pfam | PF06507 | 1.2E-37 | 287 | 369 | IPR010525 | Auxin response factor |
| Pfam | PF02309 | 1.4E-11 | 742 | 850 | IPR033389 | AUX/IAA domain |
| SuperFamily | SSF54277 | 2.94E-12 | 758 | 836 | No hit | No description |
| PROSITE profile | PS51745 | 26.82 | 759 | 843 | IPR000270 | PB1 domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0008285 | Biological Process | negative regulation of cell proliferation | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009911 | Biological Process | positive regulation of flower development | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0010227 | Biological Process | floral organ abscission | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0048481 | Biological Process | plant ovule development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 873 aa Download sequence Send to blast |
MPDLGANNGG SNQSNSSNGC VDGYSSKPNE GGLVLAQIGG SHGYQNSSPS AHDALYEELW 60 HACAGPLVTL PRIGERVFYF PQGHMEQVEA STNQGADQHM PLFNLPYKIL CRVINVQLKA 120 ELDTDEVFSQ ITLVPEAEQD ESSVDKEPLP PLPPKPLVYS FCKTLTASDT STHGGFSVLR 180 RHADECLPPL DMSQQPPSQE LVAKDLHGVE WRFRHIFRGQ PRRHLLTTGW SVFVSSKRLV 240 AGDAFIFLRG ENGELRVGVR RAMRQQNNVP SSVISSHSMH LGVIATASHA VSTRTMFSVY 300 YKPRTSPSEF IIPYDQYMKA MKINFSVGMR FKMKFEGEEV PEQRFTGTIV GICDVDPVSW 360 RDSKWRCLKV QWDEISAIAR PERVSPWKLE SSLTPAAVNS LPVARGKRPR PNILPSSSDL 420 SALDKASVDS TQVHRFPRVL QGQEVMTLGG SLGDSDLESC QKMVVWGSKL DDVKAEGMGC 480 QRRLVSENWM PPLRHDSLYS DAFPSFQPMG EVPEYRGSLT NSILEDGQQS RLSRKQFQDQ 540 EGKIVDASGL WSTSFPNSLQ EMCESNRKMS VTSAAQSHNQ SGNVRWTGLN GHPQFSNIGM 600 EASVGNWPMH MISPCQSEVS GAHGGVDRNA ARTLQVSNTG ATEQQPSFDL WEMSKGAEVV 660 SRQIDSNCKL FGFHLIDNST GGELNPSKIR GSVNEEELQG AAHASTEILS QFVEPDQQSE 720 QTKTAKSDVP AGSCEQEKTS PRSSQETQYR ASAQSNPTRS CTKVHKQGSA LGRAVDLTKF 780 EGYTDLIREL EVMFNFEGQL QDASKGWQVV YTDDEGDMML VGDDPWQEFC SMVRKIFIYT 840 REEVEKMTPR APPNMKLPGH SEEEKVARET PTR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 4ldv_A | 1e-179 | 53 | 390 | 17 | 353 | Auxin response factor 1 |
| 4ldw_A | 1e-179 | 53 | 390 | 17 | 353 | Auxin response factor 1 |
| 4ldw_B | 1e-179 | 53 | 390 | 17 | 353 | Auxin response factor 1 |
| Search in ModeBase | ||||||
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 406 | 410 | KRPRP |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Auxin response factors (ARFs) are transcriptional factors that bind specifically to the DNA sequence 5'-TGTCTC-3' found in the auxin-responsive promoter elements (AuxREs). Could act as transcriptional activator or repressor. Formation of heterodimers with Aux/IAA proteins may alter their ability to modulate early auxin response genes expression. Promotes flowering, stamen development, floral organ abscission and fruit dehiscence. Functions independently of ethylene and cytokinin response pathways. May act as a repressor of cell division and organ growth. {ECO:0000269|PubMed:12036261, ECO:0000269|PubMed:15960614, ECO:0000269|PubMed:16176952, ECO:0000269|PubMed:16339187}. | |||||
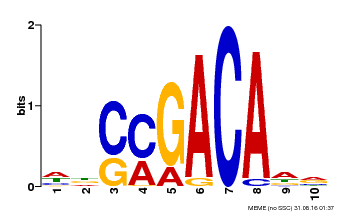
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00574 | DAP | Transfer from AT5G62000 | Download |
| |||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT116582 | 0.0 | BT116582.1 Picea glauca clone GQ03802_H23 mRNA sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_010942378.1 | 0.0 | auxin response factor 23 | ||||
| Swissprot | Q94JM3 | 0.0 | ARFB_ARATH; Auxin response factor 2 | ||||
| TrEMBL | A0A0C9RYI4 | 0.0 | A0A0C9RYI4_9SPER; Auxin response factor | ||||
| STRING | Aquca_100_00036.1 | 0.0 | (Aquilegia coerulea) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP550 | 15 | 82 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G62000.3 | 0.0 | auxin response factor 2 | ||||



