 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | MA_10428446g0010 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Acrogymnospermae; Pinidae; Pinales; Pinaceae; Picea
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 440aa MW: 48674.8 Da PI: 6.0315 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 29 | 1.8e-09 | 369 | 401 | 23 | 55 |
SS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 23 rypsaeereeLAkklgLterqVkvWFqNrRake 55
+yps++++ LA+++gL+++q+ +WF N+R ++
MA_10428446g0010 369 PYPSESQKIALAESTGLDQKQINNWFINQRKRH 401
8*****************************885 PP
| |||||||
| 2 | ELK | 40.4 | 6e-14 | 321 | 342 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK+qLlrKYsgyL+sLkqEF+
MA_10428446g0010 321 ELKDQLLRKYSGYLSSLKQEFL 342
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 2.1E-18 | 177 | 221 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 1.7E-21 | 178 | 220 | IPR005540 | KNOX1 |
| SMART | SM01256 | 3.4E-24 | 226 | 277 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 4.4E-23 | 229 | 276 | IPR005541 | KNOX2 |
| PROSITE profile | PS51213 | 11.592 | 321 | 341 | IPR005539 | ELK domain |
| Pfam | PF03789 | 7.2E-11 | 321 | 342 | IPR005539 | ELK domain |
| SMART | SM01188 | 7.1E-8 | 321 | 342 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.6 | 341 | 404 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 4.4E-13 | 343 | 408 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 6.84E-19 | 343 | 417 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 5.4E-27 | 346 | 406 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 2.55E-12 | 353 | 405 | No hit | No description |
| Pfam | PF05920 | 1.7E-16 | 361 | 400 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 379 | 402 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0001708 | Biological Process | cell fate specification | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0010051 | Biological Process | xylem and phloem pattern formation | ||||
| GO:0010089 | Biological Process | xylem development | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 440 aa Download sequence Send to blast |
MMALMTEHQP ESESIMTSIP IPSSYSSYSH GHGHGHGHGH GHGQSECLLS AAAMFQSSRP 60 CQGANKFEPA NAHIGGGGAG HFVMEQFIPE QAVISDSSIS SVKTEVCSGS GGTGVPGQFE 120 LIRRKEEGRC ARAYAEPSFV VTPLVTSLPP QQQEARMVTS LAVDMDSSCS CKPIEADAMK 180 AKIIAHVHYP RLVAAYIDCQ KVGAPPDVVS ELDELSQKCH AQQCVATISI GADPELDQFM 240 EAYCEMFIKY QEELTKPFKE AMAFLKKIEN QLGALTKGTI RTSSLDQGDE RGDGAASSEE 300 EDGSGGEVEF HEVDPHAEDR ELKDQLLRKY SGYLSSLKQE FLKKKKKGKL PKEARQKLLD 360 WWTRNYKWPY PSESQKIALA ESTGLDQKQI NNWFINQRKR HWKPSEEMQF VVMDSPNPHN 420 AAFFLEGHLR TDGTAFSMDC |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | May play a role in meristem function, and may be involved in maintaining cells in an undifferentiated, meristematic state, and its expression disappears at the same time the shoot apex undergoes the transition from vegetative to reproductive development (PubMed:11934861). Positive regulator of LATERAL ORGAN BOUNDARIES (LOB) (PubMed:11934861). Probably binds to the DNA sequence 5'-TGAC-3' (PubMed:11934861). Able to traffic from the L1 to the L2/L3 layers of the meristem, presumably through plasmodesmata (PubMed:12900451). {ECO:0000269|PubMed:11934861, ECO:0000269|PubMed:12900451, ECO:0000269|PubMed:7866029}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00675 | ChIP-seq | Transfer from GRMZM2G017087 | Download |
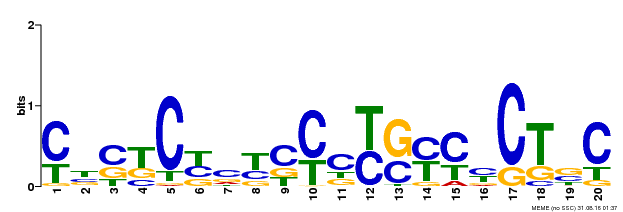
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Negatively regulated by ASYMMETRIC LEAVES1 (AS1) and ASYMMETRIC LEAVES2 (AS2). | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AF483277 | 0.0 | AF483277.1 Picea abies KNOTTED1-like homeodomain protein 2 (HBK2) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022870905.1 | 1e-108 | homeotic protein knotted-1 isoform X1 | ||||
| Swissprot | P46639 | 1e-108 | KNAT1_ARATH; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | Q8GZN0 | 0.0 | Q8GZN0_PICAB; KNOTTED1-like homeodomain protein 2 (Fragment) | ||||
| STRING | Cagra.7524s0001.1.p | 1e-107 | (Capsella grandiflora) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Representative plant | OGRP167 | 17 | 148 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G08150.1 | 1e-98 | KNOTTED-like from Arabidopsis thaliana | ||||



