 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Oropetium_20150105_22902A | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Cynodonteae; Tripogoninae; Oropetium
|
||||||||
| Family | TALE | ||||||||
| Protein Properties | Length: 274aa MW: 30353.3 Da PI: 5.4449 | ||||||||
| Description | TALE family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 27.6 | 4.9e-09 | 230 | 262 | 23 | 55 |
SS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHH CS
Homeobox 23 rypsaeereeLAkklgLterqVkvWFqNrRake 55
+yp++e++ LA+ +gL+ +q+ +WF N+R ++
Oropetium_20150105_22902A 230 PYPTEEDKVTLASMTGLDPKQINNWFINQRKRH 262
8*****************************885 PP
| |||||||
| 2 | ELK | 34.4 | 4.8e-12 | 182 | 203 | 1 | 22 |
ELK 1 ELKhqLlrKYsgyLgsLkqEFs 22
ELK++Ll+KYsg+L+ L++EF+
Oropetium_20150105_22902A 182 ELKDMLLKKYSGCLSRLRSEFL 203
9********************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM01255 | 5.6E-22 | 37 | 81 | IPR005540 | KNOX1 |
| Pfam | PF03790 | 6.5E-21 | 39 | 78 | IPR005540 | KNOX1 |
| SMART | SM01256 | 3.1E-27 | 86 | 137 | IPR005541 | KNOX2 |
| Pfam | PF03791 | 7.9E-24 | 90 | 136 | IPR005541 | KNOX2 |
| PROSITE profile | PS51213 | 10.441 | 182 | 202 | IPR005539 | ELK domain |
| Pfam | PF03789 | 5.4E-9 | 182 | 203 | IPR005539 | ELK domain |
| SMART | SM01188 | 2.0E-6 | 182 | 203 | IPR005539 | ELK domain |
| PROSITE profile | PS50071 | 12.228 | 202 | 265 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.6E-13 | 204 | 269 | IPR001356 | Homeobox domain |
| SuperFamily | SSF46689 | 4.71E-19 | 204 | 268 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-27 | 207 | 267 | IPR009057 | Homeodomain-like |
| CDD | cd00086 | 1.77E-12 | 214 | 266 | No hit | No description |
| Pfam | PF05920 | 1.2E-17 | 222 | 261 | IPR008422 | Homeobox KN domain |
| PROSITE pattern | PS00027 | 0 | 240 | 263 | IPR017970 | Homeobox, conserved site |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009736 | Biological Process | cytokinin-activated signaling pathway | ||||
| GO:0010094 | Biological Process | specification of carpel identity | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 274 aa Download sequence Send to blast |
MEDLYSIHPG IPRVGGVVMS EASVVGVGGP SPADLTELMK AQIASHPRYP SLLSAYIECR 60 KVGAPPEVAS LLEEISRDRR AGGGSGAGEI GVDPELDEFM DTYCRVLVRY KEELSRPFDE 120 AASFLSSIQT QLSNLCSAGS SPVATATHSD EMMCSSEDEQ CSGDTDAADI GQEQSSRLAD 180 HELKDMLLKK YSGCLSRLRS EFLKKRKKGK LPKDARLVLL DWWNAHYRWP YPTEEDKVTL 240 ASMTGLDPKQ INNWFINQRK RHWKPSEDMR SVL* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 197 | 206 | LRSEFLKKRK |
| 2 | 203 | 207 | KKRKK |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor that may be involved in shoot formation during early embryogenesis. {ECO:0000269|PubMed:10488233}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00670 | PBM | Transfer from GRMZM2G087741 | Download |
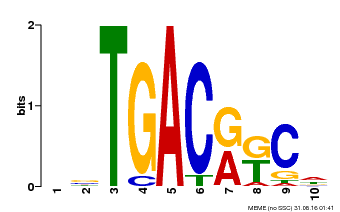
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Oropetium_20150105_22902A |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002455513.1 | 1e-171 | homeobox protein knotted-1-like 1 | ||||
| Swissprot | Q9FP29 | 1e-140 | KNOS1_ORYSJ; Homeobox protein knotted-1-like 1 | ||||
| TrEMBL | C5XI60 | 1e-170 | C5XI60_SORBI; Uncharacterized protein | ||||
| STRING | Sb03g012480.1 | 1e-171 | (Sorghum bicolor) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1698 | 38 | 92 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G23380.1 | 3e-82 | KNOTTED1-like homeobox gene 6 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Oropetium_20150105_22902A |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



