 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Oropetium_20150105_18649A | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Cynodonteae; Tripogoninae; Oropetium
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 528aa MW: 57281.6 Da PI: 4.8948 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 52.5 | 1.2e-16 | 37 | 84 | 1 | 48 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHHT CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqkyl 48
+g+WT Ed lvd+vk++G g+W+++ + g+ R++k+c++rw ++l
Oropetium_20150105_18649A 37 KGPWTSAEDAVLVDYVKKHGEGNWNAVQKNTGLSRCGKSCRLRWANHL 84
79******************************************9986 PP
| |||||||
| 2 | Myb_DNA-binding | 52.8 | 9.3e-17 | 90 | 133 | 1 | 46 |
TSSS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHH CS
Myb_DNA-binding 1 rgrWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqk 46
+g++T+eE+ l++++++++G++ W++ a++++ gRt++++k++w++
Oropetium_20150105_18649A 90 KGAFTPEEERLIIQLHAKMGNK-WARMAAHLP-GRTDNEIKNYWNT 133
799*******************.*********.***********96 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 17.289 | 32 | 84 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.42E-31 | 34 | 131 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.6E-14 | 36 | 86 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.6E-14 | 37 | 84 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 4.5E-24 | 38 | 91 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 9.87E-11 | 39 | 84 | No hit | No description |
| PROSITE profile | PS51294 | 25.656 | 85 | 139 | IPR017930 | Myb domain |
| SMART | SM00717 | 2.1E-17 | 89 | 137 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 6.5E-16 | 90 | 133 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 2.9E-26 | 92 | 137 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.43E-13 | 92 | 133 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009789 | Biological Process | positive regulation of abscisic acid-activated signaling pathway | ||||
| GO:0043068 | Biological Process | positive regulation of programmed cell death | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0045926 | Biological Process | negative regulation of growth | ||||
| GO:0048235 | Biological Process | pollen sperm cell differentiation | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 528 aa Download sequence Send to blast |
MYRLKSEDDC EMVLQDQMDS PVADDGGSPH KGPPLKKGPW TSAEDAVLVD YVKKHGEGNW 60 NAVQKNTGLS RCGKSCRLRW ANHLRPNLKK GAFTPEEERL IIQLHAKMGN KWARMAAHLP 120 GRTDNEIKNY WNTRIKRCQR AGLPVYPASV CHQSPNEDEQ VSDDFNCSEN RASDFMNGDG 180 LLLPDFTSDN VIPDALSYAP QLSAASISNL LGHSFASRSC SFMDQVDQAG ILKQSGSVLP 240 ALSDTVDGVL SSVDHFSNDS EMLKQALGFD YLNEANACSK TAAPFGVALS GSHAFLNGTF 300 SASRPINGPL KMELPSLQDT ESDPNSWLKY AVVPAMQPTE LVDPYLQSAA VTQSVKSECA 360 SPRNSGLLEE LLHEAQVLRS GNNQQLSVRS SSSSAGTPGE TTTAVSPEFD MGQEYWEEHR 420 GSFLNEYTPF SGYSFTESTA PVSAASLQLS KVSPVQSPSM GSGDQVTEHK YKPGGSPHPE 480 NIRPDAFFSG NTADVSIFND AIAMLLGNDI SSTWSNMPHA CEMSEFH* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1a5j_A | 9e-33 | 33 | 137 | 3 | 106 | B-MYB |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator of gibberellin-dependent alpha-amylase expression in aleurone cells. Involved in pollen and floral organs development. May bind to the 5'-TAACAAA-3' box of alpha-amylase promoter. {ECO:0000269|PubMed:9150608}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00490 | DAP | Transfer from AT5G06100 | Download |
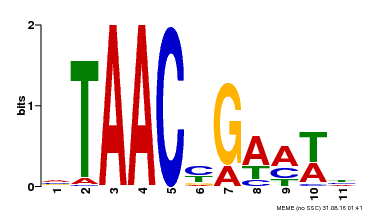
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Oropetium_20150105_18649A |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By gibberellin in aleurone cells. {ECO:0000269|PubMed:9150608}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025819835.1 | 0.0 | transcription factor GAMYB-like | ||||
| Swissprot | A2WW87 | 0.0 | GAM1_ORYSI; Transcription factor GAMYB | ||||
| TrEMBL | A0A2S3HQQ9 | 0.0 | A0A2S3HQQ9_9POAL; Transcription factor | ||||
| TrEMBL | A0A2T7DFC7 | 0.0 | A0A2T7DFC7_9POAL; Transcription factor | ||||
| STRING | Pavir.Ea03206.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3947 | 36 | 64 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G06100.3 | 1e-89 | myb domain protein 33 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Oropetium_20150105_18649A |



