 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Oropetium_20150105_16555A | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Cynodonteae; Tripogoninae; Oropetium
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 449aa MW: 48672.6 Da PI: 7.0783 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 172.1 | 1.7e-53 | 33 | 162 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLpkkvka.eekewyfFskrdkkyatgkrknratksgyW 83
lppGfrFhPtdeelvv+yLkkk+++ +l++ ++i+evd+yk++Pw+Lp+k++ +e+ewyfFs+rd+ky++g r+nra++sgyW
Oropetium_20150105_16555A 33 LPPGFRFHPTDEELVVHYLKKKAASMPLPV-TIIAEVDLYKFDPWELPEKASFgTEQEWYFFSPRDRKYPNGARPNRAATSGYW 115
79****************************.89***************544432677*************************** PP
NAM 84 katgkdkevlsk..kgelvglkktLvfykgrapkgektdWvmheyrl 128
katg+dk+++++ ++e+vg+kk Lvfy+g+ pkg kt+W+mheyrl
Oropetium_20150105_16555A 116 KATGTDKPIMASggSREKVGVKKALVFYRGKPPKGLKTNWIMHEYRL 162
***********9777788***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 6.15E-60 | 26 | 172 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 61.475 | 33 | 233 | IPR003441 | NAC domain |
| Pfam | PF02365 | 1.0E-27 | 34 | 162 | IPR003441 | NAC domain |
| SuperFamily | SSF101941 | 6.15E-60 | 221 | 233 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 449 aa Download sequence Send to blast |
MMRSMESTDS SSGSVPPQRT QQPGPVAVPA PYLPPGFRFH PTDEELVVHY LKKKAASMPL 60 PVTIIAEVDL YKFDPWELPE KASFGTEQEW YFFSPRDRKY PNGARPNRAA TSGYWKATGT 120 DKPIMASGGS REKVGVKKAL VFYRGKPPKG LKTNWIMHEY RLTDAASXXX XXXXXXXXXX 180 XXXXXXXXXX XXXXXSLIYI GIDWLNYWPG VTDRVLVSVS QLDDWVLCRI YKKINKHGVG 240 DQLRNMECGG EGSVEDATVS AYQSHAAAVA GMTLGGANYT SLLHHHHEDN FLDGLLTTVE 300 DGGGLSAAGA ASLSQLAAVT RASNTAVTKQ LLAPPAAATP FNWLDASSSG IAILPPAKRY 360 HGFSRDNADG STSLSSPAAE GNQLPAGSID GAGGSGTAAI PTTFLNPXXX XXXXXXXXXX 420 XXXXXXXXLP PETAFQQPYQ LSGVNWNP* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 2e-63 | 33 | 233 | 17 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 2e-63 | 33 | 233 | 17 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 2e-63 | 33 | 233 | 17 | 165 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 2e-63 | 33 | 233 | 17 | 165 | NO APICAL MERISTEM PROTEIN |
| 4dul_A | 2e-63 | 33 | 233 | 17 | 165 | NAC domain-containing protein 19 |
| 4dul_B | 2e-63 | 33 | 233 | 17 | 165 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor of the NAC family associated with the grain protein content (GPC). Accelerates senescence and increases nutrient remobilization from leaves to developing grains. Sequences of 11 European varieties of H.vulgare tested belongs to the same haplotype while the sequence found in H.spontaneum, an ancestor of the cultivated H.vulgare which has a higher GPC, belongs to an other haplotype. {ECO:0000269|PubMed:17124321}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00058 | PBM | Transfer from AT1G61110 | Download |
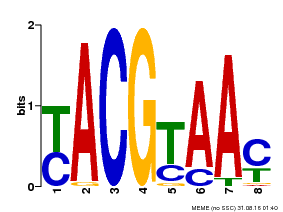
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Oropetium_20150105_16555A |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | BT085989 | 1e-143 | BT085989.1 Zea mays full-length cDNA clone ZM_BFc0095M07 mRNA, complete cds. | |||
| GenBank | BT086467 | 1e-143 | BT086467.1 Zea mays full-length cDNA clone ZM_BFc0169O24 mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_025802635.1 | 1e-154 | NAC transcription factor NAM-B2-like | ||||
| Swissprot | A0SPJ9 | 1e-142 | NAM2_HORVV; NAC transcription factor NAM-2 | ||||
| TrEMBL | A0A3L6E021 | 1e-157 | A0A3L6E021_MAIZE; NAC transcription factor ONAC010 | ||||
| STRING | GRMZM2G179885_P02 | 1e-151 | (Zea mays) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP7572 | 36 | 49 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G61110.1 | 4e-80 | NAC domain containing protein 25 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Oropetium_20150105_16555A |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



