 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Oropetium_20150105_00219A | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; PACMAD clade; Chloridoideae; Cynodonteae; Tripogoninae; Oropetium
|
||||||||
| Family | HD-ZIP | ||||||||
| Protein Properties | Length: 337aa MW: 36816.1 Da PI: 6.4534 | ||||||||
| Description | HD-ZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Homeobox | 55.7 | 8.2e-18 | 112 | 165 | 3 | 56 |
--SS--HHHHHHHHHHHHHSSS--HHHHHHHHHHCTS-HHHHHHHHHHHHHHHH CS
Homeobox 3 kRttftkeqleeLeelFeknrypsaeereeLAkklgLterqVkvWFqNrRakek 56
k+++++ eq+++Le+ Fe +++ e++ +LA+ lgL+ rqV +WFqNrRa++k
Oropetium_20150105_00219A 112 KKRRLNVEQVRTLEKNFELGNKLEPERKLQLARALGLQPRQVAIWFQNRRARWK 165
4568999**********************************************9 PP
| |||||||
| 2 | HD-ZIP_I/II | 127.8 | 4.5e-41 | 111 | 201 | 1 | 91 |
HD-ZIP_I/II 1 ekkrrlskeqvklLEesFeeeekLeperKvelareLglqprqvavWFqnrRARtktkqlEkdyeaLkraydalkeenerLekev 84
ekkrrl+ eqv++LE++Fe +kLeperK +lar+Lglqprqva+WFqnrRAR+ktkqlEkdy++Lkr++da+k++n++L +++
Oropetium_20150105_00219A 111 EKKRRLNVEQVRTLEKNFELGNKLEPERKLQLARALGLQPRQVAIWFQNRRARWKTKQLEKDYDVLKRQLDAVKADNDALLSHN 194
69********************************************************************************** PP
HD-ZIP_I/II 85 eeLreel 91
++L++e+
Oropetium_20150105_00219A 195 KKLQAEI 201
****987 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Gene3D | G3DSA:1.10.10.60 | 1.9E-18 | 92 | 168 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.97E-19 | 103 | 169 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS50071 | 17.346 | 107 | 167 | IPR001356 | Homeobox domain |
| SMART | SM00389 | 1.3E-17 | 110 | 171 | IPR001356 | Homeobox domain |
| CDD | cd00086 | 7.00E-17 | 112 | 168 | No hit | No description |
| Pfam | PF00046 | 3.3E-15 | 112 | 165 | IPR001356 | Homeobox domain |
| PRINTS | PR00031 | 1.6E-5 | 138 | 147 | IPR000047 | Helix-turn-helix motif |
| PROSITE pattern | PS00027 | 0 | 142 | 165 | IPR017970 | Homeobox, conserved site |
| PRINTS | PR00031 | 1.6E-5 | 147 | 163 | IPR000047 | Helix-turn-helix motif |
| Pfam | PF02183 | 1.9E-14 | 167 | 206 | IPR003106 | Leucine zipper, homeobox-associated |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009744 | Biological Process | response to sucrose | ||||
| GO:0048826 | Biological Process | cotyledon morphogenesis | ||||
| GO:0080022 | Biological Process | primary root development | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 337 aa Download sequence Send to blast |
MRPMASNGMA SSPSSFFPPN FLLQMQQTPP DHDSQEHHLL PPPPLHPHHN PFLPSSQQCP 60 SLQDFRGMSP MLGKRPIYGP DVVGGDELNG DGGGGGANED ELSDDGSQAG EKKRRLNVEQ 120 VRTLEKNFEL GNKLEPERKL QLARALGLQP RQVAIWFQNR RARWKTKQLE KDYDVLKRQL 180 DAVKADNDAL LSHNKKLQAE ILALKGGREA GSEQINLNKE TEASCSNRSE NSSEINLDIS 240 RTPPTESPMD PPAHHQHAGN GCGGVGMIPF YPSVGRPTGI GIDQLLHSTS GPKMEQHGGN 300 GGGCVQAPET ASFGNLLCGV DEQPPFWPWA DHQHFH* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 159 | 167 | RRARWKTKQ |
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Probable transcription factor. {ECO:0000250}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
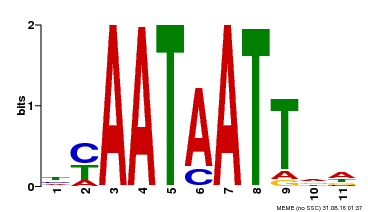
| Motif ID | Method | Source | Motif file |
| MP00225 | DAP | Transfer from AT1G69780 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | Oropetium_20150105_00219A |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | EU966490 | 1e-170 | EU966490.1 Zea mays clone 294849 homeobox-leucine zipper protein HAT7 mRNA, complete cds. | |||
| GenBank | KJ728506 | 1e-170 | KJ728506.1 Zea mays clone pUT6811 HB transcription factor (HB102) mRNA, partial cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_002465782.1 | 0.0 | homeobox-leucine zipper protein HOX21 | ||||
| Swissprot | Q8S7W9 | 1e-144 | HOX21_ORYSJ; Homeobox-leucine zipper protein HOX21 | ||||
| TrEMBL | A0A3L6SDB5 | 0.0 | A0A3L6SDB5_PANMI; Uncharacterized protein | ||||
| STRING | Pavir.Ia04323.1.p | 0.0 | (Panicum virgatum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2289 | 38 | 94 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G69780.1 | 7e-80 | HD-ZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | Oropetium_20150105_00219A |



