 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ote238018650001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Ocimeae; Ocimum
|
||||||||
| Family | MIKC_MADS | ||||||||
| Protein Properties | Length: 242aa MW: 27268.1 Da PI: 8.6058 | ||||||||
| Description | MIKC_MADS family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | SRF-TF | 84.6 | 6e-27 | 9 | 58 | 1 | 50 |
S---SHHHHHHHHHHHHHHHHHHHHHHHHHHT-EEEEEEE-TTSEEEEEE CS
SRF-TF 1 krienksnrqvtfskRrngilKKAeELSvLCdaevaviifsstgklyeys 50
k+ien rqvtfskRr+g++KKA+EL vLC+aevaviifs+tgkl+e++
Ote238018650001|238018650001 9 KKIENVNSRQVTFSKRRAGLFKKANELAVLCEAEVAVIIFSNTGKLFEFA 58
68***********************************************8 PP
| |||||||
| 2 | K-box | 61.8 | 2.8e-21 | 84 | 165 | 19 | 100 |
K-box 19 qelakLkkeienLqreqRhllGedLesLslkeLqqLeqqLekslkkiRskKnellleqieelqkkekelqeenkaLrkkle 99
+e++ Lk+eie+L+ +q +llG+dL+ ++ +eL+ Le+qL ++l i+++K++lll+++ + + +e+++ en++Lrk++e
Ote238018650001|238018650001 84 KEADLLKEEIEKLKLKQLRLLGKDLTGMTSQELHLLERQLSEGLMCIKERKEQLLLQELTQCKIQEQRAALENETLRKQVE 164
799****************************************************************************98 PP
K-box 100 e 100
e
Ote238018650001|238018650001 165 E 165
7 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS50066 | 31.413 | 1 | 61 | IPR002100 | Transcription factor, MADS-box |
| SMART | SM00432 | 1.1E-39 | 1 | 60 | IPR002100 | Transcription factor, MADS-box |
| CDD | cd00265 | 1.62E-36 | 2 | 67 | No hit | No description |
| SuperFamily | SSF55455 | 4.97E-29 | 2 | 75 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.5E-27 | 3 | 23 | IPR002100 | Transcription factor, MADS-box |
| PROSITE pattern | PS00350 | 0 | 3 | 57 | IPR002100 | Transcription factor, MADS-box |
| Pfam | PF00319 | 1.2E-25 | 10 | 57 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.5E-27 | 23 | 38 | IPR002100 | Transcription factor, MADS-box |
| PRINTS | PR00404 | 1.5E-27 | 38 | 59 | IPR002100 | Transcription factor, MADS-box |
| PROSITE profile | PS51297 | 11.515 | 79 | 169 | IPR002487 | Transcription factor, K-box |
| Pfam | PF01486 | 1.2E-14 | 84 | 163 | IPR002487 | Transcription factor, K-box |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010047 | Biological Process | fruit dehiscence | ||||
| GO:0010262 | Biological Process | somatic embryogenesis | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0048577 | Biological Process | negative regulation of short-day photoperiodism, flowering | ||||
| GO:0060862 | Biological Process | negative regulation of floral organ abscission | ||||
| GO:0060867 | Biological Process | fruit abscission | ||||
| GO:0071365 | Biological Process | cellular response to auxin stimulus | ||||
| GO:2000692 | Biological Process | negative regulation of seed maturation | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0042803 | Molecular Function | protein homodimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 242 aa Download sequence Send to blast |
MGRGKIEIKK IENVNSRQVT FSKRRAGLFK KANELAVLCE AEVAVIIFSN TGKLFEFANS 60 RYKKYVETAK GVAGEPQPEP LQPKEADLLK EEIEKLKLKQ LRLLGKDLTG MTSQELHLLE 120 RQLSEGLMCI KERKEQLLLQ ELTQCKIQEQ RAALENETLR KQVEELRTFF PPSTSPTPLC 180 IDYHSSVQKS NTAVKEGSRS PDTVADEDLD TTLQLELGLS YGSSRKRKAS DGETHSSTST 240 RN |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5f28_A | 9e-18 | 1 | 85 | 1 | 92 | MEF2C |
| 5f28_B | 9e-18 | 1 | 85 | 1 | 92 | MEF2C |
| 5f28_C | 9e-18 | 1 | 85 | 1 | 92 | MEF2C |
| 5f28_D | 9e-18 | 1 | 85 | 1 | 92 | MEF2C |
| Search in ModeBase | ||||||
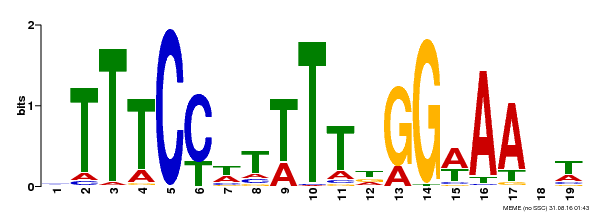
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00508 | DAP | Transfer from AT5G13790 | Download |
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011091774.1 | 1e-114 | agamous-like MADS-box protein AGL15 | ||||
| TrEMBL | A0A067L5A5 | 9e-89 | A0A067L5A5_JATCU; Uncharacterized protein | ||||
| STRING | Migut.N01802.1.p | 2e-87 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA8775 | 22 | 26 |



