 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ote100140760001 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Ocimeae; Ocimum
|
||||||||
| Family | bHLH | ||||||||
| Protein Properties | Length: 588aa MW: 65534.3 Da PI: 6.7028 | ||||||||
| Description | bHLH family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | HLH | 39.6 | 9.6e-13 | 423 | 469 | 4 | 55 |
HHHHHHHHHHHHHHHHHHHHHCTSCC.C...TTS-STCHHHHHHHHHHHHHHH CS
HLH 4 ahnerErrRRdriNsafeeLrellPk.askapskKlsKaeiLekAveYIksLq 55
+h e+Er+RR+++N++f Lr ++P+ + K++Ka+ L A+ YI +Lq
Ote100140760001|100140760001 423 NHVEAERQRREKLNQRFYALRAVVPNiS------KMDKASLLGDAIAYITELQ 469
799***********************66......*****************99 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF14215 | 8.0E-42 | 79 | 234 | IPR025610 | Transcription factor MYC/MYB N-terminal |
| PROSITE profile | PS50888 | 16.85 | 419 | 468 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SuperFamily | SSF47459 | 2.09E-18 | 422 | 486 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd00083 | 1.12E-13 | 422 | 473 | No hit | No description |
| Gene3D | G3DSA:4.10.280.10 | 2.6E-18 | 423 | 485 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| Pfam | PF00010 | 3.3E-10 | 423 | 469 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| SMART | SM00353 | 1.3E-16 | 425 | 474 | IPR011598 | Myc-type, basic helix-loop-helix (bHLH) domain |
| CDD | cd04873 | 0.0092 | 523 | 571 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| GO:0046983 | Molecular Function | protein dimerization activity | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 588 aa Download sequence Send to blast |
MGIVDWSNED KAMAAAVLGT KAFDYLMSSS VSAECSLMAM GNDENLQNKL ADLVERPNSX 60 XXXXXXXXXX XXXXXXXXEL VLGWGDGCCR EVREDEESEV TRILKMRLED ESQQTLRKRV 120 LQRLHTLFGG GDDVNYAFGL DKVTDTEMFF LASMYFSFPR GEGGPGKCFG SSKHVWLSDA 180 LKASADYCVR SYLAKSAGMQ TIVLIPTDVG VVELGSVRCI PESLELVNMV ESSFLSFSSL 240 LRSKQTAAVA AVTDKKDTNG PIPNFAVGKI FGQDLNSSHV QFRENVSVRK GEVHNRTWEA 300 SGKNMNRLPF TSNRNGFHGA AWTQYSNVKL GNPVEVYSPQ TPNKSQNREE FVLNKFQHQK 360 PAQMQIDFSG ATPRPLNSHQ QNLESELSDV EASCKEEVAG LSEDKRPRKR GRKPSNGRDE 420 PLNHVEAERQ RREKLNQRFY ALRAVVPNIS KMDKASLLGD AIAYITELQK KLKDMESDRE 480 SISREASGSE AVSTREMQDL ASCVKIEAGD EEVTVRVSCL LDDHPASRVF RAINDVQATI 540 VEAKMATGSE RVFHTFVIKS HGSHMLTKEK LIEAFSCKPN SVHRLSLA |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5gnj_A | 1e-28 | 417 | 482 | 2 | 67 | Transcription factor MYC2 |
| 5gnj_B | 1e-28 | 417 | 482 | 2 | 67 | Transcription factor MYC2 |
| 5gnj_E | 1e-28 | 417 | 482 | 2 | 67 | Transcription factor MYC2 |
| 5gnj_F | 1e-28 | 417 | 482 | 2 | 67 | Transcription factor MYC2 |
| 5gnj_G | 1e-28 | 417 | 482 | 2 | 67 | Transcription factor MYC2 |
| 5gnj_I | 1e-28 | 417 | 482 | 2 | 67 | Transcription factor MYC2 |
| 5gnj_M | 1e-28 | 417 | 482 | 2 | 67 | Transcription factor MYC2 |
| 5gnj_N | 1e-28 | 417 | 482 | 2 | 67 | Transcription factor MYC2 |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that negatively regulates jasmonate (JA) signaling (PubMed:30610166). Negatively regulates JA-dependent response to wounding, JA-induced expression of defense genes, JA-dependent responses against herbivorous insects, and JA-dependent resistance against Botrytis cinerea infection (PubMed:30610166). Plays a positive role in resistance against the bacterial pathogen Pseudomonas syringae pv tomato DC3000 (PubMed:30610166). {ECO:0000269|PubMed:30610166}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00100 | PBM | Transfer from AT1G01260 | Download |
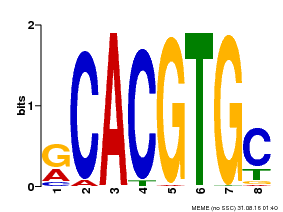
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by wounding, feeding with herbivorous insects, infection with the fungal pathogen Botrytis cinerea and infection with the bacterial pathogen Pseudomonas syringae pv tomato DC3000. {ECO:0000269|PubMed:30610166}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011093944.1 | 0.0 | transcription factor bHLH13-like | ||||
| Swissprot | A0A3Q7ELQ2 | 0.0 | MTB1_SOLLC; Transcription factor MTB1 | ||||
| TrEMBL | A0A2Z4K512 | 0.0 | A0A2Z4K512_TECGR; BHLH1 protein | ||||
| STRING | Solyc01g096050.2.1 | 0.0 | (Solanum lycopersicum) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA3585 | 23 | 48 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



