 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ote100124510011 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Ocimeae; Ocimum
|
||||||||
| Family | MYB_related | ||||||||
| Protein Properties | Length: 266aa MW: 29349.4 Da PI: 11.0936 | ||||||||
| Description | MYB_related family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 38.6 | 2.5e-12 | 60 | 104 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+W++eE+ l++ + ++ G+g+W+ I+r k+Rt+ q+ s+ qky
Ote100124510011|100124510011 60 PWSEEEHRLFLLGLRKVGKGDWRGISRNHVKTRTPTQVASHAQKY 104
8**************************9999*************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 15.158 | 53 | 109 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 2.39E-16 | 55 | 109 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.7E-16 | 56 | 107 | IPR006447 | Myb domain, plants |
| SMART | SM00717 | 1.2E-6 | 57 | 107 | IPR001005 | SANT/Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 3.4E-10 | 59 | 103 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 6.3E-10 | 60 | 104 | IPR001005 | SANT/Myb domain |
| CDD | cd00167 | 5.66E-10 | 60 | 105 | No hit | No description |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009723 | Biological Process | response to ethylene | ||||
| GO:0009733 | Biological Process | response to auxin | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0009739 | Biological Process | response to gibberellin | ||||
| GO:0009751 | Biological Process | response to salicylic acid | ||||
| GO:0009753 | Biological Process | response to jasmonic acid | ||||
| GO:0031540 | Biological Process | regulation of anthocyanin biosynthetic process | ||||
| GO:0046686 | Biological Process | response to cadmium ion | ||||
| GO:0080167 | Biological Process | response to karrikin | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003682 | Molecular Function | chromatin binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 266 aa Download sequence Send to blast |
MLFGVRVLTE GSFRKSFSLN NLAQYDQLPQ DSNNDVAAGY VSDDVVHASG RSRGRKRGVP 60 WSEEEHRLFL LGLRKVGKGD WRGISRNHVK TRTPTQVASH AQKYFLHRNN HSRRRRRSSL 120 FDITNDAVFG SEADDQMYLR NISQPQATQN QTKLSKNKKF QITNSPMAIT PLDVPITAEX 180 XXXXXXPPLG QVNQLSNSLN PIRPIPISPV PPSSKMANLN LKKLTAATIE PPQHSLPARH 240 ASAFQAMPSG AAGFNGGSGD SMISVA |
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00630 | PBM | Transfer from PK17526.1 | Download |
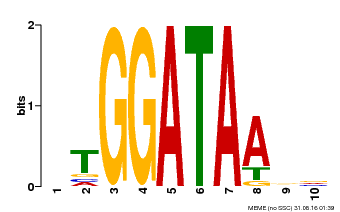
| |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011086441.1 | 1e-111 | transcription factor MYB1R1 | ||||
| TrEMBL | A0A4D9BPI4 | 1e-112 | A0A4D9BPI4_SALSN; Uncharacterized protein | ||||
| STRING | Migut.L01371.1.p | 1e-100 | (Erythranthe guttata) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA11423 | 22 | 25 |



