 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | Ote100055870051 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; eudicotyledons; Gunneridae; Pentapetalae; asterids; lamiids; Lamiales; Lamiaceae; Nepetoideae; Ocimeae; Ocimum
|
||||||||
| Family | MYB | ||||||||
| Protein Properties | Length: 257aa MW: 28312.6 Da PI: 8.0099 | ||||||||
| Description | MYB family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | Myb_DNA-binding | 34.9 | 3.5e-11 | 10 | 54 | 3 | 45 |
SS-HHHHHHHHHHHHHTTTT...-HHHHHHHHTTTS-HHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGgg...tWktIartmgkgRtlkqcksrwq 45
W+ eEd+++ +a++ ++ + +W++Ia ++ g++ +++k+++
Ote100055870051|100055870051 10 VWSREEDKQFEKAIATYPEEscdRWEKIAGEVA-GKSVEEIKHHYE 54
6*****************99*************.**********96 PP
| |||||||
| 2 | Myb_DNA-binding | 44.1 | 4.7e-14 | 113 | 157 | 3 | 47 |
SS-HHHHHHHHHHHHHTTTT-HHHHHHHHTTTS-HHHHHHHHHHH CS
Myb_DNA-binding 3 rWTteEdellvdavkqlGggtWktIartmgkgRtlkqcksrwqky 47
+WT++E+ l++ + +++G+g+W++I+r + +Rt+ q+ s+ qky
Ote100055870051|100055870051 113 AWTEDEHRLFLLGLEKYGKGDWRSISRNFVVTRTPTQVASHAQKY 157
6*******************************************9 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51293 | 9.169 | 6 | 66 | IPR017884 | SANT domain |
| SMART | SM00717 | 2.7E-11 | 7 | 59 | IPR001005 | SANT/Myb domain |
| Pfam | PF00249 | 1.4E-9 | 10 | 54 | IPR001005 | SANT/Myb domain |
| SuperFamily | SSF46689 | 5.18E-12 | 10 | 65 | IPR009057 | Homeodomain-like |
| CDD | cd00167 | 6.92E-11 | 11 | 57 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 1.2E-5 | 11 | 54 | IPR009057 | Homeodomain-like |
| PROSITE profile | PS51294 | 17.61 | 105 | 162 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 6.35E-17 | 108 | 163 | IPR009057 | Homeodomain-like |
| SMART | SM00717 | 4.5E-11 | 110 | 160 | IPR001005 | SANT/Myb domain |
| TIGRFAMs | TIGR01557 | 1.2E-17 | 110 | 160 | IPR006447 | Myb domain, plants |
| CDD | cd00167 | 7.68E-11 | 113 | 158 | No hit | No description |
| Gene3D | G3DSA:1.10.10.60 | 4.2E-11 | 113 | 156 | IPR009057 | Homeodomain-like |
| Pfam | PF00249 | 5.9E-12 | 113 | 157 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 257 aa Download sequence Send to blast |
MPGVESSSCV WSREEDKQFE KAIATYPEES CDRWEKIAGE VAGKSVEEIK HHYELLVDDV 60 SRIESGFVEV PCYNSSSSSD EDGAAKKSAN SSHLKCDGFK PSRSDQERRK GIAWTEDEHR 120 LFLLGLEKYG KGDWRSISRN FVVTRTPTQV ASHAQKYFIR LNSMNKDRRR SSIHDITSVE 180 GGDVSAPQGP ITGQSTARGP NAKSGKQSPG GSMYGHPIVG TPVNLPPPPP HMTYGVPVLP 240 GPTMNVAYPM PHTSVRR |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 2cjj_A | 2e-15 | 1 | 76 | 1 | 76 | RADIALIS |
| Search in ModeBase | ||||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that coordinates abscisic acid (ABA) biosynthesis and signaling-related genes via binding to the specific promoter motif 5'-(A/T)AACCAT-3'. Represses ABA-mediated salt (e.g. NaCl and KCl) stress tolerance. Regulates leaf shape and promotes vegetative growth. {ECO:0000269|PubMed:26243618}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00500 | DAP | Transfer from AT5G08520 | Download |
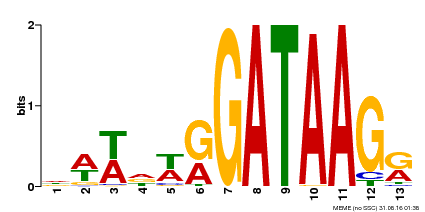
| |||
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by salicylic acid (SA) and gibberellic acid (GA) (PubMed:16463103). Triggered by dehydration and salt stress (PubMed:26243618). {ECO:0000269|PubMed:16463103, ECO:0000269|PubMed:26243618}. | |||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_011070471.1 | 1e-124 | transcription factor SRM1 | ||||
| Refseq | XP_011070472.1 | 1e-124 | transcription factor SRM1 | ||||
| Refseq | XP_020547442.1 | 1e-124 | transcription factor SRM1 | ||||
| Refseq | XP_020547443.1 | 1e-124 | transcription factor SRM1 | ||||
| Swissprot | Q9FNN6 | 1e-104 | SRM1_ARATH; Transcription factor SRM1 | ||||
| TrEMBL | A0A2G9HW37 | 1e-124 | A0A2G9HW37_9LAMI; Zuotin | ||||
| STRING | EMJ24454 | 1e-116 | (Prunus persica) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Asterids | OGEA4149 | 24 | 45 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



