 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | estExt_fgenesh1_pg.C_Chr_16.00010090 | ||||||||
| Common Name | OT_ostta16g01340 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Chlorophyta; prasinophytes; Mamiellophyceae; Mamiellales; Bathycoccaceae; Ostreococcus
|
||||||||
| Family | ARR-B | ||||||||
| Protein Properties | Length: 370aa MW: 40351 Da PI: 8.7467 | ||||||||
| Description | ARR-B family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 90 | 2.1e-28 | 212 | 265 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kpr++W+ eLH++Fv av+qL G++kA+Pk+il+lm+++gLt+e+v+SHLQkYRl
estExt_fgenesh1_pg.C_Chr_16.00010090 212 KPRVVWSAELHTQFVTAVNQL-GIDKAVPKRILDLMGIQGLTRENVASHLQKYRL 265
79*******************.********************************8 PP
| |||||||
| 2 | Response_reg | 83.1 | 8.8e-28 | 20 | 128 | 1 | 109 |
EEEESSSHHHHHHHHHHHHHTTCEEEEEESSHHHHHHHHHHHH..ESEEEEESSCTTSEHHHHHHHHHHHTTT CS
Response_reg 1 vlivdDeplvrellrqalekegyeevaeaddgeealellkekd..pDlillDiempgmdGlellkeireeepk 71
v++vdD+ l +++++++l+ +y +v++++++e+al++l++++ +D++l D+ mp++dG++ll+ i++e +
estExt_fgenesh1_pg.C_Chr_16.00010090 20 VMVVDDDLLCLKVVEKMLKACKY-RVTACSTAESALAILRTRKdeFDIVLSDVHMPDVDGFKLLEIIQFEL-N 90
89*********************.***************888888**********************9977.* PP
SEEEEEESTTTHHHHHHHHHTTESEEEESS--HHHHHH CS
Response_reg 72 lpiivvtahgeeedalealkaGakdflsKpfdpeelvk 109
lp+++++a+++++ +l+ + Ga d+l Kp+ +eel++
estExt_fgenesh1_pg.C_Chr_16.00010090 91 LPVLMMSANSDSSVVLRGIIHGAVDYLLKPVRIEELRN 128
************************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF52172 | 1.71E-33 | 17 | 139 | IPR011006 | CheY-like superfamily |
| Gene3D | G3DSA:3.40.50.2300 | 2.2E-38 | 17 | 164 | No hit | No description |
| SMART | SM00448 | 1.5E-29 | 18 | 130 | IPR001789 | Signal transduction response regulator, receiver domain |
| PROSITE profile | PS50110 | 39.97 | 19 | 134 | IPR001789 | Signal transduction response regulator, receiver domain |
| Pfam | PF00072 | 2.1E-23 | 20 | 129 | IPR001789 | Signal transduction response regulator, receiver domain |
| CDD | cd00156 | 7.59E-27 | 21 | 133 | No hit | No description |
| PROSITE profile | PS51294 | 10.846 | 209 | 268 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 1.27E-19 | 210 | 269 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 6.5E-30 | 210 | 270 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 2.6E-24 | 212 | 265 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 1.4E-7 | 214 | 264 | IPR001005 | SANT/Myb domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009873 | Biological Process | ethylene-activated signaling pathway | ||||
| GO:0010082 | Biological Process | regulation of root meristem growth | ||||
| GO:0010119 | Biological Process | regulation of stomatal movement | ||||
| GO:0010150 | Biological Process | leaf senescence | ||||
| GO:0071368 | Biological Process | cellular response to cytokinin stimulus | ||||
| GO:0080113 | Biological Process | regulation of seed growth | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 370 aa Download sequence Send to blast |
MGTAPNRLFD SKNFPAGMGV MVVDDDLLCL KVVEKMLKAC KYRVTACSTA ESALAILRTR 60 KDEFDIVLSD VHMPDVDGFK LLEIIQFELN LPVLMMSANS DSSVVLRGII HGAVDYLLKP 120 VRIEELRNIW QHVVRRDYSA RNSGSEDGVN PSSPSKRLKT SGSDSKSEEV ERAANEAGSS 180 KARKKPAGKK GAKGSKEKEA ERKGDAIDSS KKPRVVWSAE LHTQFVTAVN QLGIDKAVPK 240 RILDLMGIQG LTRENVASHL QKYRLYLKRL QGNDLMRNGS NASSSGGVSQ SRAKEQPMTI 300 KGKNVNPFLD DNIFRADDLG LPGDLGNIGY DASFDPSLLA PLDLDKVGKS DVLGADPSDN 360 ILNFFFQGD* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1irz_A | 4e-21 | 209 | 270 | 2 | 63 | ARR10-B |
| Search in ModeBase | ||||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00054 | PBM | Transfer from AT4G16110 | Download |
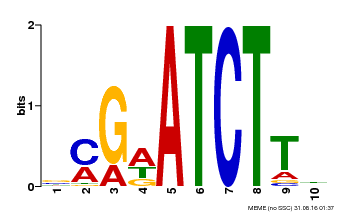
| |||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_022840333.1 | 0.0 | Signal transduction response regulator, receiver domain | ||||
| TrEMBL | A0A096P7X7 | 0.0 | A0A096P7X7_OSTTA; Signal transduction response regulator, receiver domain | ||||
| STRING | A0A096P7X7 | 0.0 | (Ostreococcus tauri) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Chlorophytae | OGCP1220 | 15 | 16 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G16110.1 | 2e-78 | response regulator 2 | ||||
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



