 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os11g45530.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | FAR1 | ||||||||
| Protein Properties | Length: 795aa MW: 92498 Da PI: 8.4152 | ||||||||
| Description | FAR1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | FAR1 | 42.8 | 1.7e-13 | 46 | 121 | 10 | 90 |
FAR1 10 vGFsvrkskskkskrngeitkrtfvCskegkreeekkktekerrtraetrtgCkaklkvkkekdgkwevtkleleHnHela 90
GF v k+ skk+ ++g++ t+ Cs++gk++ ++++ + +++++tr +C ak++ ++ d+k+++++l l+HnH ++
LOC_Os11g45530.1 46 KGFGVIKRGSKKT-EDGKVRYFTLACSRQGKAQYTSTN---KFKPNPSTRQQCPAKVNFYLH-DEKFCISTLTLDHNHVVS 121
69***99999988.8899****************9888...99******************9.9**************876 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF03101 | 7.5E-11 | 46 | 121 | IPR004330 | FAR1 DNA binding domain |
| Pfam | PF10551 | 1.9E-38 | 242 | 336 | IPR018289 | MULE transposase domain |
| PROSITE profile | PS50966 | 9.458 | 525 | 563 | IPR007527 | Zinc finger, SWIM-type |
| Pfam | PF04434 | 2.1E-4 | 537 | 558 | IPR007527 | Zinc finger, SWIM-type |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0010018 | Biological Process | far-red light signaling pathway | ||||
| GO:0042753 | Biological Process | positive regulation of circadian rhythm | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0008270 | Molecular Function | zinc ion binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 795 aa Download sequence Send to blast |
MDTNLCSDDE QEDHIISDED GFAIEEVESD SNALVADKTK LLEPTKGFGV IKRGSKKTED 60 GKVRYFTLAC SRQGKAQYTS TNKFKPNPST RQQCPAKVNF YLHDEKFCIS TLTLDHNHVV 120 SPSKARHLRC HKKLDLQAKR RLELNDQARI RINKNFSSLV MQAGGYENLQ FGEKECRNYL 180 QDVRKLKLGA GDAHAVYQYF LRMTSKDPNF FYVMDVDEDS RLKNVLWVDA RSRATYESFS 240 DVVTFDTTYL TNKYHMPFAP FVGVNHHGES VLLGCALLSN EETETFVWLF RSWLSCMSNK 300 APNAIITDQC RAMQNAIMEV FPEARHRWCL WHIMKKIPEK LGGYLEYEVI SSTLSNVVYD 360 SLNRDDFDKG WMKMIDEFSL QDNEWLAGLY DNRELWVPAY VKDTFWAGMS SSQRSESVNA 420 FFDGYVNART TLKQFVEQYD NALRDKVEKE NKSDCKSFQE AIPCITHYEF ERQFQVAYTN 480 KKFKEFQDEL RGKIYYYATL RKTEGLVHTF SVREDRKIGE QRVVSELLVL FNQEDCDLHC 540 ECRHFEFRGI LCRHILSILP LVDIEKVPSK YILQRWRKDF KRKHTFIKCS YDDQLDTPIV 600 RRFDTLCKRF NEVAENGSVS DALCNLVMDG LNELQIKIDA HHGSKEIQEY QQNGKNKDMV 660 LKQGKMVLSP ISVRRRGRPP SLRKQSKLDQ VVRRLRMKKQ QESTIEVQSR RRRKTRTKNV 720 ISKDKQLINI QNSRQMEVNF DHCYGGAYEA GSAEFAANEF QGLQSNHPAP PSHISSYMDL 780 LQVSYKRRTH ASDV* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 785 | 789 | KRRTH |
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| Expression Atlas | Q2QZM0 | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
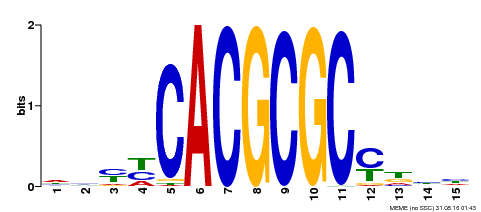
| Motif ID | Method | Source | Motif file |
| MP00434 | DAP | Transfer from AT4G15090 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os11g45530.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | - | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os11g45530 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AC136998 | 0.0 | AC136998.2 Oryza sativa (japonica cultivar-group) chromosome 11 BAC clone OSJNBa0082P17, complete sequence. | |||
| GenBank | AP014967 | 0.0 | AP014967.1 Oryza sativa Japonica Group DNA, chromosome 11, cultivar: Nipponbare, complete sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_024314597.1 | 0.0 | protein FAR1-RELATED SEQUENCE 6-like | ||||
| TrEMBL | Q2QZM0 | 0.0 | Q2QZM0_ORYSJ; Transposon protein, putative, unclassified | ||||
| STRING | OS11T0681300-00 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP18 | 32 | 967 | Representative plant | OGRP30 | 12 | 351 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT4G15090.1 | 1e-127 | FAR1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os11g45530.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



