 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os07g39220.1 | ||||||||
| Common Name | BZR1, LOC4343719, Os07g0580500, OsJ_24880, P0453G03.22 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | BES1 | ||||||||
| Protein Properties | Length: 299aa MW: 31908.7 Da PI: 8.6003 | ||||||||
| Description | BES1 family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | DUF822 | 221.7 | 1.5e-68 | 6 | 147 | 1 | 145 |
DUF822 1 ggsgrkptwkErEnnkrRERrRRaiaakiyaGLRaqGnyklpkraDnneVlkALcreAGwvvedDGttyrkgskpleeaeaagssasaspess 93
+++gr+ptwkErEnnkrRERrRRaiaaki++GLRa Gny+lpk++DnneVlkALcreAGwvvedDGttyrkg+kp+ ++a+g+s +sp+ss
LOC_Os07g39220.1 6 AAAGRTPTWKERENNKRRERRRRAIAAKIFTGLRALGNYNLPKHCDNNEVLKALCREAGWVVEDDGTTYRKGCKPP-PSSAGGASVGMSPCSS 97
589*************************************************************************.9999************ PP
DUF822 94 lq.sslkssalaspvesysaspksssfpspssldsislasaasllpvlsvlsl 145
+q +s+ ss+++spv+sy+asp+sssfpsps++d+ s+ ++llp+l+ l++
LOC_Os07g39220.1 98 TQlLSAPSSSFPSPVPSYHASPASSSFPSPSRIDNPSA---SCLLPFLRGLPN 147
*********************************99866...588888876655 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF05687 | 2.9E-64 | 8 | 132 | IPR008540 | BES1/BZR1 plant transcription factor, N-terminal |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009741 | Biological Process | response to brassinosteroid | ||||
| GO:0009742 | Biological Process | brassinosteroid mediated signaling pathway | ||||
| GO:0040008 | Biological Process | regulation of growth | ||||
| GO:0042742 | Biological Process | defense response to bacterium | ||||
| GO:0045892 | Biological Process | negative regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0005737 | Cellular Component | cytoplasm | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0001046 | Molecular Function | core promoter sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0005515 | Molecular Function | protein binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009030 | anatomy | carpel | ||||
| PO:0007130 | developmental stage | sporophyte reproductive stage | ||||
| PO:0007606 | developmental stage | gynoecium development stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 299 aa Download sequence Send to blast |
MTSGAAAAGR TPTWKERENN KRRERRRRAI AAKIFTGLRA LGNYNLPKHC DNNEVLKALC 60 REAGWVVEDD GTTYRKGCKP PPSSAGGASV GMSPCSSTQL LSAPSSSFPS PVPSYHASPA 120 SSSFPSPSRI DNPSASCLLP FLRGLPNLPP LRVSSSAPVT PPLSSPTASR PPKIRKPDWD 180 VDPFRHPFFA VSAPASPTRG RRLEHPDTIP ECDESDVSTV DSGRWISFQM ATTAPTSPTY 240 NLVNPGASTS NSMEIEGTAG RGGAEFEFDK GRVTPWEGER IHEVAAEELE LTLGVGAK* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 5zd4_A | 3e-28 | 10 | 89 | 372 | 451 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_B | 3e-28 | 10 | 89 | 372 | 451 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_C | 3e-28 | 10 | 89 | 372 | 451 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| 5zd4_D | 3e-28 | 10 | 89 | 372 | 451 | Maltose-binding periplasmic protein,Protein BRASSINAZOLE-RESISTANT 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.54781 | 0.0 | callus| flower| leaf| panicle| root | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32991957 | 0.0 | ||||
| Expression Atlas | Q7XI96 | - | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Positive brassinosteroid-signaling protein. Mediates downstream brassinosteroid-regulated growth response and feedback inhibition of brassinosteroid biosynthetic genes. May act as transcriptional repressor by binding the brassinosteroid-response element (5'-CGTGCG-3') in the promoter of GRAS32 (AC Q9LWU9), another positive regulator of brassinosteroid signaling (By similarity). {ECO:0000250}. | |||||
| UniProt | Positive brassinosteroid-signaling protein. Mediates downstream brassinosteroid-regulated growth response and feedback inhibition of brassinosteroid (BR) biosynthetic genes (PubMed:17699623, PubMed:19220793). May act as transcriptional repressor by binding the brassinosteroid-response element (BREE) (5'-CGTG(T/C)G-3') in the promoter of DLT (AC Q9LWU9), another positive regulator of BR signaling (PubMed:19220793). Acts as transcriptional repressor of LIC, a negative regulator of BR signaling, by binding to the BRRE element of its promoter. BZR1 and LIC play opposite roles in BR signaling and regulation of leaf bending (PubMed:22570626). {ECO:0000269|PubMed:17699623, ECO:0000269|PubMed:19220793, ECO:0000269|PubMed:22570626}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
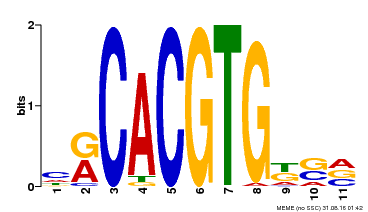
| Motif ID | Method | Source | Motif file |
| MP00073 | ChIP-chip | Transfer from AT1G19350 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os07g39220.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Down-regulated by 24-epibrassinolide. {ECO:0000269|PubMed:22570626}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os07g39220 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK106748 | 0.0 | AK106748.1 Oryza sativa Japonica Group cDNA clone:002-115-C06, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015645123.1 | 0.0 | protein BZR1 homolog 1 | ||||
| Swissprot | B8B7S5 | 0.0 | BZR1_ORYSI; Protein BZR1 homolog 1 | ||||
| Swissprot | Q7XI96 | 0.0 | BZR1_ORYSJ; Protein BZR1 homolog 1 | ||||
| TrEMBL | A0A0E0QAC1 | 1e-173 | A0A0E0QAC1_ORYRU; Uncharacterized protein | ||||
| TrEMBL | I1QEM4 | 1e-173 | I1QEM4_ORYGL; Uncharacterized protein | ||||
| STRING | OS07T0580500-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP5474 | 34 | 53 | Representative plant | OGRP3719 | 10 | 24 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT1G19350.3 | 2e-80 | BES1 family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os07g39220.1 |
| Entrez Gene | 4343719 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



