 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os06g49040.1 | ||||||||
| Common Name | LOC4341990, OJ1215_E11.18, Os06g0703900, OsJ_22555, OSNPB_060703900 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 480aa MW: 52205 Da PI: 6.0972 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 101.8 | 4.2e-32 | 269 | 323 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
k+rlrWt eLHerFveav++L G+ekAtPk +l+lmkv+gLt++hvkSHLQkYRl
LOC_Os06g49040.1 269 KARLRWTLELHERFVEAVNKLDGPEKATPKGVLKLMKVEGLTIYHVKSHLQKYRL 323
68****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 11.393 | 266 | 326 | IPR017930 | Myb domain |
| SuperFamily | SSF46689 | 7.88E-17 | 267 | 323 | IPR009057 | Homeodomain-like |
| Gene3D | G3DSA:1.10.10.60 | 2.4E-30 | 267 | 324 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 1.1E-24 | 269 | 324 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 3.9E-8 | 271 | 322 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 3.6E-22 | 359 | 404 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0020039 | anatomy | leaf lamina | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 480 aa Download sequence Send to blast |
MSSQSVVAVK QITAPDKIVE TCPSTKHSAH KLFDVKPDFQ GLIDDNLSSS SQSSSIKIEL 60 IRSSSLPNIL PFQKRSSEPE PESPLSHVSH PNVSEPVYSN SSTFCTSLFS SSSMETEPCR 120 QLGTLPFLPH PPKCEQQVSA GHSSSSSLLV PGGDGDIGNA HDEPEQSDDL KDFLNLSGGD 180 ASDGSFHGEN NAMAFAEQME FQFLSEQLGI AITDNEESPR LDDIYGTPPQ LSSLPVSSCS 240 NQSVQKAGSP VKVQLSSPRS SSGSATTNKA RLRWTLELHE RFVEAVNKLD GPEKATPKGV 300 LKLMKVEGLT IYHVKSHLQK YRLAKYLPET KEDKKASSED KKSQSGSSGN DSVKKKNLQV 360 AEALRMQMEV QKQLHEQLEV QRQLQLRIEE HARYLQRILE EQHKVSISSN SLSLKPPAES 420 QPESPKPTSE KKEAESEAGA ATSAQPSSED KSPDAECKSS PPVGSKRARV HVGDEHQCS* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4k_A | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4k_B | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_A | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_C | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_D | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_F | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_H | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j5b_J | 8e-26 | 269 | 326 | 2 | 59 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.5279 | 0.0 | flower| leaf| panicle| seed| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32977968 | 0.0 | ||||
| Expression Atlas | Q5Z811 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | DEVELOPMENTAL STAGE: Expressed throughout all stages of plant growth. Increased expression in leaves before the booting stage. {ECO:0000269|PubMed:26082401}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the root cap and in the exodermis of the root, in the root tip of lateral roots, in the mesophyll cells of the leaf, in pollen, vascular cylinder of the anther and the veins of the lemma, palea and pistils, and in the xylem and phloem regions of large vascular bundles, small vascular bundles and diffuse vascular bundles in node I (PubMed:26082401). {ECO:0000269|PubMed:26082401}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor involved in phosphate starvation signaling (PubMed:26082401). Binds to P1BS, an imperfect palindromic sequence 5'-GNATATNC-3', to promote the expression of inorganic phosphate (Pi) starvation-responsive genes (PubMed:26082401). Functionally redundant with PHR1 and PHR2 in regulating Pi starvation response and Pi homeostasis (PubMed:26082401). {ECO:0000269|PubMed:26082401}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00354 | DAP | Transfer from AT3G13040 | Download |
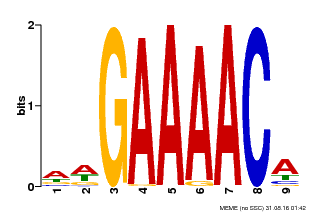
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os06g49040.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated under Pi starvation conditions. {ECO:0000269|PubMed:26082401}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os06g49040 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK060648 | 0.0 | AK060648.1 Oryza sativa Japonica Group cDNA clone:001-027-G12, full insert sequence. | |||
| GenBank | AK067950 | 0.0 | AK067950.1 Oryza sativa Japonica Group cDNA clone:J013127L15, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015644151.1 | 0.0 | protein PHOSPHATE STARVATION RESPONSE 3-like | ||||
| Swissprot | Q6YXZ4 | 1e-168 | PHR3_ORYSJ; Protein PHOSPHATE STARVATION RESPONSE 3 | ||||
| TrEMBL | Q5Z811 | 0.0 | Q5Z811_ORYSJ; Os06g0703900 protein | ||||
| STRING | OS06T0703900-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP4526 | 37 | 69 | Representative plant | OGRP78 | 17 | 262 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G13040.2 | 1e-66 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os06g49040.1 |
| Entrez Gene | 4341990 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



