 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os05g49420.1 | ||||||||
| Common Name | LOC4339653, OJ1781_H11.25, Os05g0569300, OSJNBa0040E06.7, OSNPB_050569300 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 381aa MW: 40408.5 Da PI: 9.6898 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 68.8 | 8.8e-22 | 245 | 307 | 1 | 63 |
XXXXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 1 ekelkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkkevaklksev 63
e+elkre+rkq+NRe+ArrsR+RK+ae+e L++ v++L+aeN++L++e+++l++ ++kl+ e+
LOC_Os05g49420.1 245 ERELKREKRKQSNRESARRSRLRKQAETEDLATQVESLTAENTSLRSEISRLSESSEKLRLEN 307
89*********************************************************9887 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| Pfam | PF07777 | 6.6E-31 | 1 | 93 | IPR012900 | G-box binding protein, multifunctional mosaic region |
| Gene3D | G3DSA:1.20.5.170 | 6.4E-17 | 242 | 304 | No hit | No description |
| SMART | SM00338 | 8.4E-22 | 245 | 309 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 2.8E-20 | 245 | 307 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 13.196 | 247 | 310 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 2.94E-11 | 248 | 303 | No hit | No description |
| CDD | cd14702 | 1.03E-22 | 250 | 298 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 252 | 267 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0009737 | Biological Process | response to abscisic acid | ||||
| GO:0005829 | Cellular Component | cytosol | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| GO:0044212 | Molecular Function | transcription regulatory region DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009066 | anatomy | anther | ||||
| PO:0001004 | developmental stage | anther development stage | ||||
| PO:0007130 | developmental stage | sporophyte reproductive stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 381 aa Download sequence Send to blast |
MAHDEAVATQ KIGKTTSPPK DQPTPCPFPD WSAVQAYYGP GVLPPTYFAP AIASGHAPPP 60 YMWGPQPIMP PPFGTPYAAM YPHGGAYPHP LMPMMANPLS MEPAKSASSK EKGSNKKLKE 120 VDGAAVSTGS GDSKKTMTSS GDYSAEGSSD VNDLKVGKTG RKRRLDDGAG AETSAAAKME 180 NALPPSHILG STAILPNHSF PAQVIRPSAT NVANSRALGT PISPPPGVIV PSHTGVSTEL 240 LIKDERELKR EKRKQSNRES ARRSRLRKQA ETEDLATQVE SLTAENTSLR SEISRLSESS 300 EKLRLENSAL MGKLKDPAAS TQAETSLQKT TTASSPRVVE NFLSMIDNTN KTSVRHTEHA 360 EPKLRQLLGS GPATDVVAAS * |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 261 | 267 | RRSRLRK |
| 2 | 261 | 268 | RRSRLRKQ |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.19553 | 0.0 | callus| flower| leaf| panicle| root| seed| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32975458 | 0.0 | ||||
| Expression Atlas | Q6AUN3 | - | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00318 | DAP | Transfer from AT2G46270 | Download |
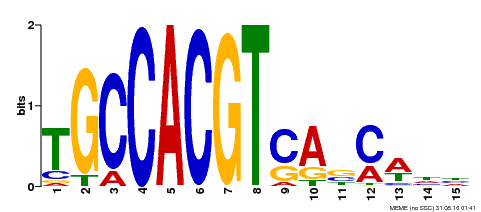
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os05g49420.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os05g49420 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK065440 | 0.0 | AK065440.1 Oryza sativa Japonica Group cDNA clone:J013020K21, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015640745.1 | 0.0 | common plant regulatory factor 1 | ||||
| TrEMBL | A0A0E0PRE9 | 0.0 | A0A0E0PRE9_ORYRU; Uncharacterized protein | ||||
| TrEMBL | Q6AUN3 | 0.0 | Q6AUN3_ORYSJ; Os05g0569300 protein | ||||
| STRING | ORUFI05G28170.1 | 0.0 | (Oryza rufipogon) | ||||
| STRING | OS05T0569300-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2776 | 38 | 88 | Representative plant | OGRP4346 | 11 | 23 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G46270.1 | 1e-45 | G-box binding factor 3 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os05g49420.1 |
| Entrez Gene | 4339653 |



