 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os05g36160.1 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 268aa MW: 27629.9 Da PI: 9.5703 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 43.8 | 5.5e-14 | 186 | 237 | 3 | 54 |
XXCHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 3 elkrerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelkk 54
+ +r++r++kNRe+A rsR+RK+a+i+eLe v+eLe+e +L e ee ++
LOC_Os05g36160.1 186 AMQRQKRMIKNRESAARSRERKQAYIAELEAQVAELEEEHAQLLREQEEKNQ 237
689**************************************99977666544 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 4.9E-9 | 184 | 249 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 10.875 | 186 | 234 | IPR004827 | Basic-leucine zipper domain |
| Pfam | PF00170 | 5.4E-12 | 186 | 237 | IPR004827 | Basic-leucine zipper domain |
| CDD | cd14707 | 6.81E-17 | 188 | 234 | No hit | No description |
| Gene3D | G3DSA:1.20.5.170 | 2.6E-13 | 189 | 235 | No hit | No description |
| SuperFamily | SSF57959 | 6.25E-11 | 189 | 235 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 191 | 206 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| GO:0043565 | Molecular Function | sequence-specific DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 268 aa Download sequence Send to blast |
MASSRVMAAA AASSSSSPPP PPPAAAAAGG AADLARFRST SSGIGSMNMD DILRNIYGEA 60 APPPGAAGSA PAPPPAGEAA GAPVAEVAAR RTAEEVWKEI SSSGGLSAPA PAPAAGAAGR 120 GGGPEMTLED FLAREDDPRA TAVEGNMVVG FPNVTEGVGT AGGGRGGGGG GRGRKRTLMD 180 PADRAAMQRQ KRMIKNRESA ARSRERKQAY IAELEAQVAE LEEEHAQLLR EQEEKNQKRL 240 KEIKEQAVAV VIRKKTQDLR RTNSMEW* |
| Nucleic Localization Signal ? help Back to Top | |||
|---|---|---|---|
| No. | Start | End | Sequence |
| 1 | 162 | 170 | GGRGGGGGG |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.37925 | 0.0 | callus| leaf| panicle| seed | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 37990279 | 0.0 | ||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription activator that binds to the ABA-responsive elements (ABREs) in vitro. Involved in abiotic stress responses and abscisic acid (ABA) signaling (PubMed:20039193). Involved in the signaling pathway that induces growth inhibition in response to D-allose (PubMed:23397192). {ECO:0000269|PubMed:20039193, ECO:0000269|PubMed:23397192}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00648 | PBM | 25215497 | Download |
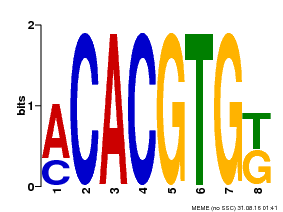
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os05g36160.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by anoxia, drought, salt stress, oxidative stress, cold and abscisic acid (ABA) (PubMed:20039193). Induced by D-allose (PubMed:23397192). {ECO:0000269|PubMed:20039193, ECO:0000269|PubMed:23397192}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | - | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os05g36160 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | CT834109 | 0.0 | CT834109.1 Oryza sativa (indica cultivar-group) cDNA clone:OSIGCSA060C07, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015639395.1 | 0.0 | bZIP transcription factor 12-like | ||||
| Swissprot | Q0JHF1 | 8e-63 | BZP12_ORYSJ; bZIP transcription factor 12 | ||||
| TrEMBL | A0A0E0PN11 | 0.0 | A0A0E0PN11_ORYRU; Uncharacterized protein | ||||
| STRING | OS05T0437700-01 | 1e-167 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP2665 | 37 | 86 | Representative plant | OGRP9845 | 4 | 10 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G44080.1 | 5e-15 | bZIP family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os05g36160.1 |
| Publications ? help Back to Top | |||
|---|---|---|---|
|
|||



