 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os05g35170.4 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 329aa MW: 36600.4 Da PI: 4.9982 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 158.3 | 3.2e-49 | 6 | 133 | 1 | 128 |
NAM 1 lppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.k.kvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdke 91
lppGfrFhPtd elv++yLk+k+ gkk + ++i++v++yk+ PwdLp + ++++++ ew+fF++rdkky++g+r+nr+t +gyWk++gkd++
LOC_Os05g35170.4 6 LPPGFRFHPTDVELVSYYLKRKIMGKKPLI-QAISDVELYKFAPWDLPaQsCLQSRDLEWFFFCPRDKKYPNGSRTNRSTPNGYWKTSGKDRT 97
79*************************777.88**************95347778888*********************************** PP
NAM 92 vlskkgelvglkktLvfykgrapkgektdWvmheyrl 128
+ +++ vg kktL+f++g+apkg++tdWvm ey++
LOC_Os05g35170.4 98 IEL-NSRIVGSKKTLIFHEGKAPKGNRTDWVMYEYKM 133
***.999****************************97 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 2.62E-58 | 3 | 156 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 56.218 | 6 | 156 | IPR003441 | NAC domain |
| Pfam | PF02365 | 6.1E-26 | 7 | 132 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 329 aa Download sequence Send to blast |
MAQTCLPPGF RFHPTDVELV SYYLKRKIMG KKPLIQAISD VELYKFAPWD LPAQSCLQSR 60 DLEWFFFCPR DKKYPNGSRT NRSTPNGYWK TSGKDRTIEL NSRIVGSKKT LIFHEGKAPK 120 GNRTDWVMYE YKMEDNQLVS AGFSKDDFVL CKIFKKSGLG PRIGEQYGAP FNEEEWEHAD 180 AEMFPLLPNV ETSVFPLLPS SEVVNSTDDT RVQPSVAARA IEELPVQHLP HVCAGNGSTY 240 QNITVTGESA LMELPSQHSV ESIGDEVVSV DNCSNVVNNA DSPVIEGLVL EELSRFLTDS 300 PHHGNPVGEV DRACSLSDLP VFLLCLGI* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 3ulx_A | 2e-51 | 2 | 157 | 11 | 169 | Stress-induced transcription factor NAC1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.12020 | 0.0 | callus| flower| leaf| panicle| root| seed| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32984749 | 0.0 | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in root meristem (Ref.1). Expressed in roots, rosette leaves, cauline leaves, shoot apex, stems and flowers (PubMed:17158162). {ECO:0000269|PubMed:17158162, ECO:0000269|Ref.1}. | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in the epidermal cells of the root apical region. {ECO:0000269|PubMed:20388856}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator activated by proteolytic cleavage through regulated intramembrane proteolysis (RIP) (By similarity). Transcripition activator associated with the induction of genes related to flavonoid biosynthesis and required for the accumulation of anthocyanins in response to high light stress (PubMed:19887540). Plays a role in the regulation of 20S and 26S proteasomes in response to high light stress (PubMed:21889048). {ECO:0000250|UniProtKB:Q949N0, ECO:0000269|PubMed:19887540, ECO:0000269|PubMed:21889048}. | |||||
| UniProt | Transcriptional regulator that binds specific DNA sequences on the promoter regions of target genes. {ECO:0000250|UniProtKB:Q9C8W9}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00583 | DAP | Transfer from AT5G64060 | Download |
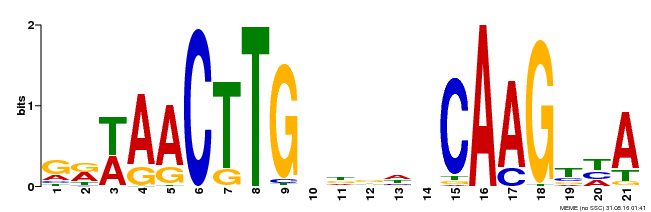
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os05g35170.4 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: By exposure to high light (PubMed:19887540). Induced by heat and methyl methanesulfonate (MMS) treatment (PubMed:17158162). {ECO:0000269|PubMed:17158162, ECO:0000269|PubMed:19887540}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os05g35170 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AB362160 | 0.0 | AB362160.1 Oryza sativa Japonica Group mRNA for transcription factor, complete cds. | |||
| GenBank | AK099540 | 0.0 | AK099540.1 Oryza sativa Japonica Group cDNA clone:J013032G17, full insert sequence. | |||
| GenBank | EU846998 | 0.0 | EU846998.1 Oryza sativa Japonica Group clone KCS104B01 NAM-like protein mRNA, complete cds. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015639254.1 | 0.0 | NAC domain-containing protein 78 isoform X1 | ||||
| Swissprot | Q84K00 | 1e-73 | NAC78_ARATH; NAC domain-containing protein 78 | ||||
| Swissprot | Q9FY82 | 2e-74 | NAC82_ARATH; NAC domain-containing protein 82 | ||||
| TrEMBL | A0A0E0HEQ9 | 0.0 | A0A0E0HEQ9_ORYNI; Uncharacterized protein | ||||
| TrEMBL | B8AYH2 | 0.0 | B8AYH2_ORYSI; Uncharacterized protein | ||||
| TrEMBL | I1PVT8 | 0.0 | I1PVT8_ORYGL; Uncharacterized protein | ||||
| TrEMBL | Q60EC0 | 0.0 | Q60EC0_ORYSJ; NAM-like protein | ||||
| STRING | OS05T0426200-02 | 0.0 | (Oryza sativa) | ||||
| STRING | ONIVA05G17930.1 | 0.0 | (Oryza nivara) | ||||
| STRING | ORGLA05G0148600.1 | 0.0 | (Oryza glaberrima) | ||||
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G64060.1 | 4e-81 | NAC domain containing protein 103 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os05g35170.4 |



