 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os03g56580.1 | ||||||||
| Common Name | LOC4334294, Os03g0777000, OsJ_12786, OSJNBa0010I09.1, OSJNBa0070N04.8, OSNPB_030777000 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | NAC | ||||||||
| Protein Properties | Length: 363aa MW: 40558.6 Da PI: 6.8387 | ||||||||
| Description | NAC family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | NAM | 163.6 | 7e-51 | 39 | 170 | 2 | 128 |
NAM 2 ppGfrFhPtdeelvveyLkkkvegkkleleevikevdiykvePwdLp.kkvkaeekewyfFskrdkkyatgkrknratksgyWkatgkdkevl 93
pGfrFhPtdeelv++yL++kv++k+l++ e+ike+diyk++PwdLp +++ ++ekewyfF+ r +ky+++ r+nr+t sg+Wkatg d++++
LOC_Os03g56580.1 39 LPGFRFHPTDEELVTFYLRRKVARKSLSI-EIIKEMDIYKHDPWDLPnASTVGGEKEWYFFCLRGRKYRNSIRPNRVTGSGFWKATGIDRPIY 130
69***************************.89***************556666899************************************* PP
NAM 94 sk.....kgelvglkktLvfykgrapkgektdWvmheyrl 128
s+ +ge +glkk+Lv+y+g+a kg+ktdW+mhe+rl
LOC_Os03g56580.1 131 SAavnsnSGESIGLKKSLVYYRGSAGKGTKTDWMMHEFRL 170
*9888888888***************************98 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SuperFamily | SSF101941 | 1.15E-55 | 38 | 197 | IPR003441 | NAC domain |
| PROSITE profile | PS51005 | 54.759 | 38 | 197 | IPR003441 | NAC domain |
| Pfam | PF02365 | 7.9E-25 | 40 | 170 | IPR003441 | NAC domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0005992 | Biological Process | trehalose biosynthetic process | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0006561 | Biological Process | proline biosynthetic process | ||||
| GO:0009718 | Biological Process | anthocyanin-containing compound biosynthetic process | ||||
| GO:0010120 | Biological Process | camalexin biosynthetic process | ||||
| GO:0042538 | Biological Process | hyperosmotic salinity response | ||||
| GO:1900056 | Biological Process | negative regulation of leaf senescence | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009005 | anatomy | root | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 363 aa Download sequence Send to blast |
MVTSKEFARD QAAMDQKIKS DVGEVVLAGD EEEDGDVVLP GFRFHPTDEE LVTFYLRRKV 60 ARKSLSIEII KEMDIYKHDP WDLPNASTVG GEKEWYFFCL RGRKYRNSIR PNRVTGSGFW 120 KATGIDRPIY SAAVNSNSGE SIGLKKSLVY YRGSAGKGTK TDWMMHEFRL PPAIAAADAS 180 PCMQEAEVWT ICRIFKRSIT YRKQQQQQAW RPPATVTVKA PPPGDSSSNT GSFESDGGGD 240 EFMNCGLTPA ISQQQQHGGR HQMMSTMSCN GGYFFNDGIH HSHSHHKLHS QWGSLQMAPP 300 EPKPEPEQKP LSSPAMTIAF HQNDHGFPAA AADFYKDGYL EEIARMMEVA DPSPTGFYDC 360 RY* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 1ut4_A | 7e-49 | 40 | 203 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut4_B | 7e-49 | 40 | 203 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_A | 7e-49 | 40 | 203 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 1ut7_B | 7e-49 | 40 | 203 | 19 | 171 | NO APICAL MERISTEM PROTEIN |
| 3swm_A | 9e-49 | 40 | 203 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_B | 9e-49 | 40 | 203 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_C | 9e-49 | 40 | 203 | 22 | 174 | NAC domain-containing protein 19 |
| 3swm_D | 9e-49 | 40 | 203 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_A | 9e-49 | 40 | 203 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_B | 9e-49 | 40 | 203 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_C | 9e-49 | 40 | 203 | 22 | 174 | NAC domain-containing protein 19 |
| 3swp_D | 9e-49 | 40 | 203 | 22 | 174 | NAC domain-containing protein 19 |
| 4dul_A | 7e-49 | 40 | 203 | 19 | 171 | NAC domain-containing protein 19 |
| 4dul_B | 7e-49 | 40 | 203 | 19 | 171 | NAC domain-containing protein 19 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.17090 | 0.0 | callus| flower| leaf | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32983562 | 0.0 | ||||
| Expression Atlas | Q8S7I7 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Expressed in roots, root caps, cotyledons, tips and margin of young leaves, senescent regions of fully expanded leaves and floral tissues, including old sepals, petals, staments, mature anthers and pollen grains. Not detected in the abscission zone of open flowers, emerging lateral roots and root meristematic zones. {ECO:0000269|PubMed:22345491}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcription factor that binds to the 5'- RRYGCCGT-3' consensus core sequence. Central longevity regulator. Negative regulator of leaf senescence. Modulates cellular H(2)O(2) levels and enhances tolerance to various abiotic stresses through the regulation of DREB2A. {ECO:0000269|PubMed:22345491}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00310 | DAP | Transfer from AT2G43000 | Download |
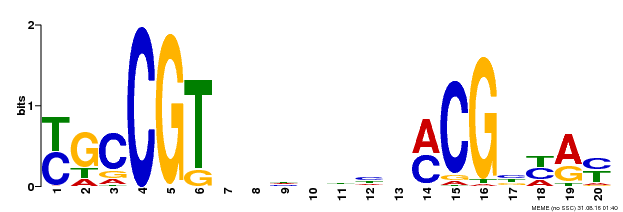
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os03g56580.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Up-regulated by H(2)O(2), paraquat, ozone, 3-aminotriazole and salt stress. {ECO:0000269|PubMed:22345491}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os03g56580 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK073539 | 0.0 | AK073539.1 Oryza sativa Japonica Group cDNA clone:J033041C24, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015628846.1 | 0.0 | transcription factor JUNGBRUNNEN 1 | ||||
| Swissprot | Q9SK55 | 6e-83 | NAC42_ARATH; Transcription factor JUNGBRUNNEN 1 | ||||
| TrEMBL | Q8S7I7 | 0.0 | Q8S7I7_ORYSJ; NAC-domain containing protein 9, putative, expressed | ||||
| STRING | OS03T0777000-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP1191 | 38 | 129 | Representative plant | OGRP17 | 15 | 800 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT2G43000.1 | 6e-84 | NAC domain containing protein 42 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os03g56580.1 |
| Entrez Gene | 4334294 |



