 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os02g52780.1 | ||||||||
| Common Name | LOC4330838, OJ1004_A11.20, Os02g0766700, OSNPB_020766700, P0539D10.39 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | bZIP | ||||||||
| Protein Properties | Length: 358aa MW: 38099.5 Da PI: 9.7632 | ||||||||
| Description | bZIP family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | bZIP_1 | 48.2 | 2.3e-15 | 277 | 325 | 5 | 53 |
CHHHCHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHHH CS
bZIP_1 5 krerrkqkNReAArrsRqRKkaeieeLeekvkeLeaeNkaLkkeleelk 53
+r+rr++kNRe+A rsRqRK+a++ eLe +v++L++ N++L+k+ +e+
LOC_Os02g52780.1 277 RRQRRMIKNRESAARSRQRKQAYMMELEAEVAKLKELNDELQKKQDEML 325
79***************************************98876643 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| SMART | SM00338 | 6.3E-14 | 273 | 338 | IPR004827 | Basic-leucine zipper domain |
| PROSITE profile | PS50217 | 11.197 | 275 | 327 | IPR004827 | Basic-leucine zipper domain |
| Gene3D | G3DSA:1.20.5.170 | 8.6E-15 | 277 | 328 | No hit | No description |
| Pfam | PF00170 | 2.6E-13 | 277 | 327 | IPR004827 | Basic-leucine zipper domain |
| SuperFamily | SSF57959 | 7.8E-11 | 277 | 327 | No hit | No description |
| CDD | cd14707 | 2.31E-24 | 277 | 329 | No hit | No description |
| PROSITE pattern | PS00036 | 0 | 280 | 295 | IPR004827 | Basic-leucine zipper domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0009414 | Biological Process | response to water deprivation | ||||
| GO:0009651 | Biological Process | response to salt stress | ||||
| GO:0009738 | Biological Process | abscisic acid-activated signaling pathway | ||||
| GO:0010255 | Biological Process | glucose mediated signaling pathway | ||||
| GO:0045893 | Biological Process | positive regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0000976 | Molecular Function | transcription regulatory region sequence-specific DNA binding | ||||
| GO:0003700 | Molecular Function | transcription factor activity, sequence-specific DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0009049 | anatomy | inflorescence | ||||
| PO:0001083 | developmental stage | inflorescence development stage | ||||
| PO:0007006 | developmental stage | IL.00 inflorescence just visible stage | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 358 aa Download sequence Send to blast |
MDFPGGSGRQ QQLPPMTPLP LARQGSVYSL TFDEFQSTLG GVGKDFGSMN MDELLRSIWT 60 AEESHAVGAA TTTTATTASV AAAEHAAVGA PPVQRQGSLT LPRTLSQKTV DEVWRDMMCF 120 GGGGASTAPA AAEPPPPAHR QQTLGEITLE EFLVRAGVVR EDMSVPPVPP APTPTAAAVP 180 PPPPPQQQTP MLFGQSNVFP PMVPPLSLGN GLVSGAVGHG GGGAASLVSP VRPVSSNGFG 240 KMEGGDLSSL SPSPVPYVFK GGLRGRKAPG IEKVVERRQR RMIKNRESAA RSRQRKQAYM 300 MELEAEVAKL KELNDELQKK QDEMLEQQKN EVLERMSRQV GPTAKRICLR RTLTGPW* |
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.11781 | 0.0 | callus| leaf| panicle| root| seed| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32982085 | 0.0 | ||||
| Expression Atlas | Q6Z312 | - | ||||
| Expression -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| Uniprot | TISSUE SPECIFICITY: Highly expressed in leaves. {ECO:0000269|PubMed:18931143}. | |||||
| Functional Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | Transcriptional activator that mediates abscisic acid (ABA) signaling (PubMed:18931143, PubMed:19947981, PubMed:27424498, PubMed:27325665). Can regulate the expression of a wide spectrum of stress-related genes in response to abiotic stresses through an ABA-dependent regulation pathway. Confers ABA-dependent drought and salinity tolerance (PubMed:18931143, PubMed:27325665). Binds specifically to the ABA-responsive elements (ABRE) in the promoter of target genes to mediate stress-responsive ABA signaling (PubMed:27325665, PubMed:27424498). {ECO:0000269|PubMed:18931143, ECO:0000269|PubMed:19947981, ECO:0000269|PubMed:27325665, ECO:0000269|PubMed:27424498}. | |||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
| Motif ID | Method | Source | Motif file |
| MP00186 | DAP | Transfer from AT1G45249 | Download |
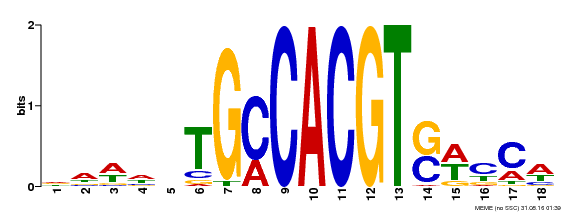
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os02g52780.1 |
| Regulation -- Description ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Description | |||||
| UniProt | INDUCTION: Induced by abscisic acid (ABA) (PubMed:18315698, PubMed:18931143). Induced by drought, salt and osmotic stresses (PubMed:18931143). {ECO:0000269|PubMed:18315698, ECO:0000269|PubMed:18931143}. | |||||
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os02g52780 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK072062 | 0.0 | AK072062.1 Oryza sativa Japonica Group cDNA clone:J013109E06, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015625853.1 | 0.0 | bZIP transcription factor 23 | ||||
| Swissprot | Q6Z312 | 0.0 | BZP23_ORYSJ; bZIP transcription factor 23 | ||||
| TrEMBL | B9F3E8 | 0.0 | B9F3E8_ORYSJ; Uncharacterized protein | ||||
| STRING | OS02T0766700-01 | 0.0 | (Oryza sativa) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP706 | 38 | 147 | Representative plant | OGRP540 | 15 | 80 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT3G19290.2 | 5e-41 | ABRE binding factor 4 | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os02g52780.1 |
| Entrez Gene | 4330838 |



