 |
PlantRegMap/PlantTFDB v5.0
Plant Transcription
Factor Database
|
| Home TFext BLAST Prediction Download Help About Links PlantRegMap |
Transcription Factor Information
| Basic Information? help Back to Top | |||||||||
|---|---|---|---|---|---|---|---|---|---|
| TF ID | LOC_Os02g47190.1 | ||||||||
| Common Name | LOC4330431, Os02g0700300, OSNPB_020700300 | ||||||||
| Organism | |||||||||
| Taxonomic ID | |||||||||
| Taxonomic Lineage |
cellular organisms; Eukaryota; Viridiplantae; Streptophyta; Streptophytina; Embryophyta; Tracheophyta; Euphyllophyta; Spermatophyta; Magnoliophyta; Mesangiospermae; Liliopsida; Petrosaviidae; commelinids; Poales; Poaceae; BOP clade; Oryzoideae; Oryzeae; Oryzinae; Oryza; Oryza sativa
|
||||||||
| Family | G2-like | ||||||||
| Protein Properties | Length: 289aa MW: 31390.5 Da PI: 8.9696 | ||||||||
| Description | G2-like family protein | ||||||||
| Gene Model |
|
||||||||
| Signature Domain? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| No. | Domain | Score | E-value | Start | End | HMM Start | HMM End |
| 1 | G2-like | 107 | 1e-33 | 31 | 85 | 1 | 55 |
G2-like 1 kprlrWtpeLHerFveaveqLGGsekAtPktilelmkvkgLtlehvkSHLQkYRl 55
kprlrWtp+LHerFveav++LGG++kAtPk++l+lm++kgLtl+h+kSHLQkYRl
LOC_Os02g47190.1 31 KPRLRWTPDLHERFVEAVTKLGGPDKATPKSVLRLMGMKGLTLYHLKSHLQKYRL 85
79****************************************************8 PP
| |||||||
| Protein Features ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Database | Entry ID | E-value | Start | End | InterPro ID | Description |
| PROSITE profile | PS51294 | 14.032 | 28 | 88 | IPR017930 | Myb domain |
| Gene3D | G3DSA:1.10.10.60 | 9.7E-33 | 28 | 86 | IPR009057 | Homeodomain-like |
| SuperFamily | SSF46689 | 1.0E-16 | 31 | 88 | IPR009057 | Homeodomain-like |
| TIGRFAMs | TIGR01557 | 9.3E-25 | 31 | 86 | IPR006447 | Myb domain, plants |
| Pfam | PF00249 | 9.0E-10 | 33 | 84 | IPR001005 | SANT/Myb domain |
| Pfam | PF14379 | 1.1E-24 | 130 | 174 | IPR025756 | MYB-CC type transcription factor, LHEQLE-containing domain |
| Gene Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| GO Term | GO Category | GO Description | ||||
| GO:0006355 | Biological Process | regulation of transcription, DNA-templated | ||||
| GO:0005634 | Cellular Component | nucleus | ||||
| GO:0003677 | Molecular Function | DNA binding | ||||
| Plant Ontology ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PO Term | PO Category | PO Description | ||||
| PO:0020039 | anatomy | leaf lamina | ||||
| Sequence ? help Back to Top |
|---|
| Protein Sequence Length: 289 aa Download sequence Send to blast |
MFEGMERAGY GVGVGGAGAV GAGVVLSRDP KPRLRWTPDL HERFVEAVTK LGGPDKATPK 60 SVLRLMGMKG LTLYHLKSHL QKYRLGKQNK KDTGLEASRG AFAAHGISFA SAAPPTIPSA 120 ENNNAGETPL ADALRYQIEV QRKLHEQLEV QKKLQMRIEA QGKYLQTILE KAQNNLSYDA 180 TGTANLEATR TQLTDFNLAL SGFMNNVSQV CEQNNGELAK AISEDNLRTT NLGFQLYHGI 240 QDSDDVKCSQ DEGLLLLDLN IKGGGYDHLS SNAMRGGESG LKISQHRR* |
| 3D Structure ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| PDB ID | Evalue | Query Start | Query End | Hit Start | Hit End | Description |
| 6j4r_A | 4e-21 | 31 | 87 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_B | 4e-21 | 31 | 87 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_C | 4e-21 | 31 | 87 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| 6j4r_D | 4e-21 | 31 | 87 | 1 | 57 | Protein PHOSPHATE STARVATION RESPONSE 1 |
| Search in ModeBase | ||||||
| Expression -- UniGene ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| UniGene ID | E-value | Expressed in | ||||
| Os.24699 | 0.0 | leaf| panicle| stem | ||||
| Expression -- Microarray ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | E-value | ||||
| GEO | 32969466 | 0.0 | ||||
| Expression Atlas | Q0DYD6 | - | ||||
| Binding Motif ? help Back to Top | |||
|---|---|---|---|
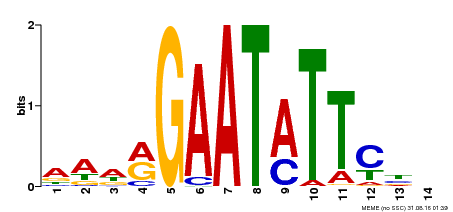
| Motif ID | Method | Source | Motif file |
| MP00544 | DAP | Transfer from AT5G45580 | Download |
| |||
| Cis-element ? help Back to Top | |
|---|---|
| Source | Link |
| PlantRegMap | LOC_Os02g47190.1 |
| Regulation -- PlantRegMap ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Upstream Regulator | Target Gene | ||||
| PlantRegMap | Retrieve | Retrieve | ||||
| Phenotype -- Mutation ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | ID | |||||
| RiceGE | Os02g47190 | |||||
| Annotation -- Nucleotide ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | |||
| GenBank | AK059448 | 0.0 | AK059448.1 Oryza sativa Japonica Group cDNA clone:001-028-A10, full insert sequence. | |||
| Annotation -- Protein ? help Back to Top | |||||||
|---|---|---|---|---|---|---|---|
| Source | Hit ID | E-value | Description | ||||
| Refseq | XP_015623757.1 | 0.0 | myb family transcription factor PHL11 | ||||
| TrEMBL | A0A0D3F9C2 | 0.0 | A0A0D3F9C2_9ORYZ; Uncharacterized protein | ||||
| TrEMBL | A0A0D9YWX9 | 0.0 | A0A0D9YWX9_9ORYZ; Uncharacterized protein | ||||
| TrEMBL | A0A0E0NJP6 | 0.0 | A0A0E0NJP6_ORYRU; Uncharacterized protein | ||||
| TrEMBL | I1P3D8 | 0.0 | I1P3D8_ORYGL; Uncharacterized protein | ||||
| TrEMBL | Q0DYD6 | 0.0 | Q0DYD6_ORYSJ; Os02g0700300 protein | ||||
| STRING | OGLUM02G29720.1 | 0.0 | (Oryza glumipatula) | ||||
| STRING | ORUFI02G30700.1 | 0.0 | (Oryza rufipogon) | ||||
| STRING | OS02T0700300-01 | 0.0 | (Oryza sativa) | ||||
| STRING | ORGLA02G0249400.1 | 0.0 | (Oryza glaberrima) | ||||
| STRING | OBART02G29100.1 | 0.0 | (Oryza barthii) | ||||
| Orthologous Group ? help Back to Top | |||
|---|---|---|---|
| Lineage | Orthologous Group ID | Taxa Number | Gene Number |
| Monocots | OGMP3940 | 34 | 70 | Representative plant | OGRP78 | 17 | 262 |
| Best hit in Arabidopsis thaliana ? help Back to Top | ||||||
|---|---|---|---|---|---|---|
| Hit ID | E-value | Description | ||||
| AT5G45580.1 | 2e-64 | G2-like family protein | ||||
| Link Out ? help Back to Top | |
|---|---|
| Phytozome | LOC_Os02g47190.1 |
| Entrez Gene | 4330431 |



